Abstract
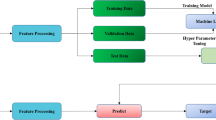
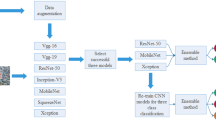
Glomerulosclerosis characterizes many conditions of primary kidney disease in advanced stages. Its accurate diagnosis relies on histological analysis of renal cortex biopsy, and it is paramount to guide the appropriate treatment and minimize the chances of the disease progressing to chronic stages. This article presents an ensemble approach composed of five convolutional neural networks (CNNs) - VGG-19, Inception-V3, ResNet-50, DenseNet-201, and EfficientNet-B2 - to detect glomerulosclerosis in glomerulus images. We fine-tuned the CNNs and evaluated several configurations for the fully connected layers. In total, we analyzed 25 different models. These CNNs, individually, demonstrated effectiveness in the task; however, we verified that the union of these five well-known CNNs improved the detection rate while decreasing the standard deviations of current techniques. The experiments were carried out in a data set comprised of 1,028 images, on which we applied data-augmentation techniques in the training set. The proposed CNNs ensemble achieved a near-perfect accuracy of 99.0% and kappa of 98.0%.









Similar content being viewed by others
References
Alves E, Souza Filho JB, Kritski AL (2019) An ensemble approach for supporting the respiratory isolation of presumed tuberculosis inpatients. Neurocomputing 331:289–300
Araújo IC, Schnitman L, Duarte AA et al (2017) Automated detection of segmental glomerulosclerosis in kidney histopathology. In: XIII Brazilian Congress on Computational Intelligence, p 12
Barros GO, Navarro B, Duarte A et al (2017) Pathospotter-k: a computational tool for the automatic identification of glomerular lesions in histological images of kidneys. Sci Rep 7(1):1–8
Barros GO, Wanderley DC, Rebouças LO et al (2022) Podnet: ensemble-based classification of podocytopathy on kidney glomerular images. In: VISIGRAPP (5: VISAPP), pp 405–412
Bevilacqua V, Pietroleonardo N, Triggiani V et al (2017) An innovative neural network framework to classify blood vessels and tubules based on haralick features evaluated in histological images of kidney biopsy. Neurocomputing 228:143–153. https://doi.org/10.1016/j.neucom.2016.09.091
Bueno G, Fernandez-Carrobles MM, Gonzalez-Lopez L et al (2020) Glomerulosclerosis identification in whole slide images using semantic segmentation. Comput Methods Programs Biomed 184(105):273
Chagas P, Souza L, Araújo I et al (2020) Classification of glomerular hypercellularity using convolutional features and support vector machine. Artif Intell Med 103(101):808
Cohen J (1960) A coefficient of agreement for nominal scales. Educ Psychol Measur 20:37–46
Dasarathy BV, Sheela BV (1979) A composite classifier system design: concepts and methodology. Proc IEEE 67(5):708–713
Dhaun N, Bellamy C, Cattran D et al (2014) Utility of renal biopsy in the clinical management of renal disease: hematuria should not be missed reply. Kidney Int 86(6):1269–1269. https://doi.org/10.1038/ki.2014.278
Dietterich TG (2000) Ensemble methods in machine learning. In: multiple classifier systems. MCS 2000. Lecture notes in computer science, vol 1857. Springer, Berlin, Heidelberg. https://doi.org/10.1007/3-540-45014-9_1
dos Santos WL, de Freitas LA, Duarte AA et al (2022) Computational pathology, new horizons and challenges for anatomical pathology. Surg Exp Pathol 5(1):1–7
Ginley B, Lutnick B, Jen KY et al (2019) Computational segmentation and classification of diabetic glomerulosclerosis. J Am Soc Nephrol 30(10):1953–1967
Ginley BG, Tomaszewski JE, Jen KY, et al (2018) Computational analysis of the structural progression of human glomeruli in diabetic nephropathy. In: Medical Imaging 2018: Digital Pathology, International Society for Optics and Photonics, p 105810A
He K, Zhang X, Ren S, et al (2015) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, Proceedings of the IEEE conference on computer vision and pattern recognition, p 770–778
Heckenauer R, Weber J, Wemmert C, et al (2020) Real-time detection of glomeruli in renal pathology. In: 2020 IEEE 33rd International Symposium on Computer-Based Medical Systems (CBMS), IEEE, pp 350–355
Hermsen M, de Bel T, Den Boer M et al (2019) Deep learning-based histopathologic assessment of kidney tissue. J Am Soc Nephrol 30(10):1968–1979
Huang G, Liu Z, Weinberger KQ (2016) Densely connected convolutional networks. CoRR abs/1608.06993. https://arxiv.org/abs/1608.06993
Jeong HJ (2020) Diagnosis of renal transplant rejection: Banff classification and beyond. Kidney Res Clin Pract 39(1):17
Kalantari A, Kamsin A, Shamshirband S et al (2018) Computational intelligence approaches for classification of medical data: state-of-the-art, future challenges and research directions. Neurocomputing 276:2–22. https://doi.org/10.1016/j.neucom.2017.01.126
Kannan S, Morgan LA, Liang B et al (2019) Segmentation of glomeruli within trichrome images using deep learning. Kidney Int Rep 4(7):955–962. https://doi.org/10.1016/j.ekir.2019.04.008
Kolachalama VB, Singh P, Lin CQ et al (2018) Association of pathological fibrosis with renal survival using deep neural networks. Kidney Int Rep 3(2):464–475. https://doi.org/10.1016/j.ekir.2017.11.002
Kornblith S, Shlens J, Le QV (2019) Do better imagenet models transfer better? In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp 2661–2671
Landis JR, Koch GG (1977) The measurement of observer agreement for categorical data. Biometrics 33(1):159–174
Marsh JN, Matlock MK, Kudose S et al (2018) Deep learning global glomerulosclerosis in transplant kidney frozen sections. IEEE Trans Med Imaging 37(12):2718–2728
Oliveira L, Chagas P, Duarte A et al (2022) PathoSpotter: computational intelligence applied to nephropathology. Springer, Cham, pp 253–272. https://doi.org/10.1007/978-3-031-11570-7_16
Piccialli F, Giampaolo F, Salvi A et al (2021) A robust ensemble technique in forecasting workload of local healthcare departments. Neurocomputing 444:69–78
Rehem JMC, dos Santos WLC, Duarte AA, et al (2021) Automatic glomerulus detection in renal histological images. In: Medical Imaging 2021: Digital Pathology, SPIE, pp 115–125
Risdon RA, Turner DR (2012) Atlas of renal pathology, vol 2. Springer, Berlin
Russakovsky O, Deng J, Su H et al (2015) Imagenet large scale visual recognition challenge. Int J Comput Vision 115:211–252
Santos JD, de MS Veras R, Silva RR et al (2021) A hybrid of deep and textural features to differentiate glomerulosclerosis and minimal change disease from glomerulus biopsy images. Biomed Signal Process Control 70(103):020. https://doi.org/10.1016/j.bspc.2021.103020
Sheehan SM, Korstanje R (2018) Automatic glomerular identification and quantification of histological phenotypes using image analysis and machine learning. Am J Physiol Renal Physiol 315(6):F1644–F1651
Shorten C, Khoshgoftaar TM (2019) A survey on image data augmentation for deep learning. J Big Data 6(1):1–48
Shubham S, Jain N, Gupta V et al (2021) Identify glomeruli in human kidney tissue images using a deep learning approach. Soft Comput. https://doi.org/10.1007/s00500-021-06143-z
Silva J, Souza L, Chagas P et al (2022) Boundary-aware glomerulus segmentation: toward one-to-many stain generalization. Comput Med Imaging Graph 100(102):104
Simonyan K, Zisserman A (2015) Very deep convolutional networks for large-scale image recognition. In: Bengio Y, LeCun Y (eds) 3rd International Conference on Learning Representations, ICLR 2015, San Diego, CA, USA, May 7-9, 2015, Conference Track Proceedings
Szegedy C, Vanhoucke V, Ioffe S, et al (2016) Rethinking the inception architecture for computer vision. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp 2818–2826
Tajbakhsh N, Shin JY, Gurudu SR et al (2016) Convolutional neural networks for medical image analysis: full training or fine tuning? IEEE Trans Med Imaging 35:1299–1312
Tan M, Le QV (2019) Efficientnet: Rethinking model scaling for convolutional neural networks. In: International Conference on Machine Learning, 2019
Tieleman T, Hinton G et al (2012) Lecture 6.5-rmsprop: divide the gradient by a running average of its recent magnitude. COURSERA Neural Networks Mach Learn 4:26–31
Van der Laak J, Litjens G, Ciompi F (2021) Deep learning in histopathology: the path to the clinic. Nat Med 27(5):775–784. https://doi.org/10.1038/s41591-021-01343-4
Yu H, Yang LT, Zhang Q et al (2021) Convolutional neural networks for medical image analysis: state-of-the-art, comparisons, improvement and perspectives. Neurocomputing 444:92–110. https://doi.org/10.1016/j.neucom.2020.04.157
Zheng Y, Cassol CA, Jung S et al (2021) Deep-learning-driven quantification of interstitial fibrosis in digitized kidney biopsies. Am J Pathol 191(8):1442–1453. https://doi.org/10.1016/j.ajpath.2021.05.005
Zheng Z, Zhang X, Ding J et al (2021) Deep learning-based artificial intelligence system for automatic assessment of glomerular pathological findings in lupus nephritis. Diagnostics. https://doi.org/10.3390/diagnostics11111983
Zhou Z (2012) Ensemble methods: foundations and algorithms. CRC Press, Boca Raton
Acknowledgments
Washington Santos and Luciano Oliveira have research scholarships from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), grants 306779/2017 and 308580/2021-4, respectively. Angelo Duarte have a research support from Universidade Estadual de Feira de Santana grant TO 074/2021.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This work was conducted in accordance with resolution No. 466/12 of the Brazilian National Health Council. To preserve confidentiality, the images (including those shown in the paper) were separated from other patient’s data. No data presented herein allows patient identification. All the procedures were approved by the Ethics Committee for Research Involving Human subjects of the Gonçalo Moniz Institute from the Oswaldo Cruz Foundation (CPqGM/FIOCRUZ), Protocols No. 188/09 and No. 1817574.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Santos, J., Silva, R., Oliveira, L. et al. Glomerulosclerosis detection with pre-trained CNNs ensemble. Comput Stat 39, 561–581 (2024). https://doi.org/10.1007/s00180-022-01307-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00180-022-01307-3