Abstract
The standard Tile Assembly Model (TAM) of Winfree (Algorithmic self-assembly of DNA, Ph.D. thesis, 1998) is a mathematical theory of crystal aggregations via monomer additions with applications to the emerging science of DNA self-assembly. Self-assembly under the rules of this model is programmable and can perform Turing universal computation. Many variations of this model have been proposed and the canonical problem of assembling squares has been studied extensively.
We consider the problem of building approximate squares in TAM. Given any \(\varepsilon \in (0,\frac{1}{4}]\) we show how to construct squares whose sides are within (1±ε)N of any given positive integer N using \(O( \frac{\log \frac{1}{\varepsilon}}{\log \log\frac{1}{\varepsilon}} + \frac{\log \log \varepsilon N}{\log \log \log \varepsilon N} )\) tile types. We prove a matching lower bound by showing that \(\varOmega( \frac{\log \frac{1}{\varepsilon}}{\log \log\frac{1}{\varepsilon}} + \frac{\log \log \varepsilon N}{\log \log \log \varepsilon N} )\) tile types are necessary almost always to build squares of required approximate dimensions. In comparison, the optimal construction for a square of side exactly N in TAM uses \(O(\frac{\log N}{\log \log N})\) tile types.
The question of constructing approximate squares has been recently studied in a modified tile assembly model involving concentration programming. All our results are trivially translated into the concentration programming model by assuming arbitrary (non-zero) concentrations for our tile types. Indeed, the non-zero concentrations could be chosen by an adversary and our results would still hold.
Our construction can get highly accurate squares using very few tile types and are feasible starting from values of N that are orders of magnitude smaller than the best comparable constructions previously suggested. At an accuracy of ε=0.01, the number of tile types used to achieve a square of size 107 is just 58 and our constructions are proven to work for all N≥13130. If the concentrations of the tile types are carefully chosen, we prove that our construction assembles an L×L square in optimal assembly time O(L) where (1−ε)N≤L≤(1+ε)N.




Similar content being viewed by others
Notes
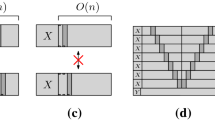
The face labels merely depict the bits of the counter and do not play any role in tile binding. Here, they are identical to the North pad type.
References
Adleman, L., Cheng, Q., Goel, A., Huang, M.D.: Running time and program size for self-assembled squares. In: Symposium on Theory of Computing, pp. 740–748 (2001)
Aggarwal, G., Cheng, Q., Goldwasser, M.H., Kao, M.Y., de Espanes, P.M., Schweller, R.T.: Complexities for generalized models of self-assembly. SIAM J. Comput. 34(6), 1493–1515 (2005)
Andersen, E., Dong, M., Nielsen, M., Jahn, K., Subramani, R., Mamdouh, W., Golas, M., Sander, B., Stark, H., Oliveira, C., Pedersen, J.S., Birkedal, V., Besenbacher, F., Gothelf, K., Kjems, J.: Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459(7243), 73–76 (2009)
Barish, R., Rothemund, P., Winfree, E.: Two computational primitives for algorithmic self-assembly: copying and counting. Nano Lett. 5, 2586–2592 (2005)
Becker, F., Rapaport, I., Remila, E.: Self-assemblying classes of shapes with a minimum number of tiles, and in optimal time. In: Foundations of Software Technology and Theoretical Computer Science, pp. 45–56 (2006)
Becker, F., Remila, E., Schabanel, N.: Time optimal self-assembling of 2D and 3D shapes: the case of squares and cubes. In: Goel, A., Simmel, F., Sosík, P. (eds.) DNA Computing. Lecture Notes in Computer Science, vol. 5347, pp. 144–155. Springer, Berlin (2009)
Berger, R.: The undecidability of the domino problem. Mem. Am. Math. Soc. 66, 1–72 (1966)
Chandran, H., Gopalkrishnan, N., Reif, J.: The tile complexity of linear assemblies. In: International Colloquium on Automata, Languages and Programming, pp. 235–253 (2009)
Chen, H.L., Goel, A.: Error free self-assembly using error prone tiles. In: Ferretti, C., Mauri, G., Zandron, C. (eds.) DNA Computing. Lecture Notes in Computer Science, vol. 3384, pp. 702–707. Springer, Berlin (2005)
Demaine, E., Demaine, M., Fekete, S., Ishaque, M., Rafalin, E., Schweller, R., Souvaine, D.: Staged self-assembly: nanomanufacture of arbitrary shapes with O(1) glues. Nat. Comput. 7(3), 347–370 (2008)
Dietz, H., Douglas, S., Shih, W.: Folding DNA into twisted and curved nanoscale shapes. Science 325(5941), 725–730 (2009)
Dirks, R., Pierce, N.: Triggered amplification by hybridization chain reaction. Proc. Natl. Acad. Sci. USA 101(43), 15275–15278 (2004)
Doty, D.: Randomized self-assembly for exact shapes. SIAM J. Comput. 39(8), 3521–3552 (2010)
Doty, D., Patitz, M., Summers, S.: Limitations of self-assembly at temperature 1. Theor. Comput. Sci. 412(1–2), 145–158 (2011)
Douglas, S., Dietz, H., Liedl, T., Hogberg, B., Graf, F., Shih, W.: Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459(7245), 414–418 (2009)
Garey, M., Johnson, D.: Computers and Intractability: A Guide to the Theory of NP-Completeness. Freeman, New York (1981)
Kao, M.Y., Schweller, R.: Reducing tile complexity for self-assembly through temperature programming. In: Symposium on Discrete Algorithms, pp. 571–580 (2006)
Kao, M.Y., Schweller, R.: Randomized self-assembly for approximate shapes. In: International Colloquium on Automata, Languages and Programming, pp. 370–384 (2008)
LaBean, T., Yan, H., Kopatsch, J., Liu, F., Winfree, E., Reif, J., Seeman, N.: Construction, analysis, ligation, and self-assembly of DNA triple crossover complexes. J. Am. Chem. Soc. 122(9), 1848–1860 (2000)
Lewis, H., Papadimitriou, C.: Elements of the Theory of Computation. Prentice Hall, New York (1981)
Mao, C., Labean, T., Reif, J., Seeman, N.: Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407, 493–496 (2000)
Park, S.H., Yin, P., Liu, Y., Reif, J., LaBean, T., Yan, H.: Programmable DNA self-assemblies for nanoscale organization of ligands and proteins. Nano Lett. 5, 729–733 (2005)
Reif, J., Sahu, S., Yin, P.: Compact error-resilient computational DNA tiling assemblies. In: Ferretti, C., Mauri, G., Zandron, C. (eds.) DNA Computing. Lecture Notes in Computer Science, vol. 3384, pp. 293–307. Springer, Berlin (2005)
Robinson, R.: Undecidability and nonperiodicity for tilings of the plane. Invent. Math. 12, 177–209 (1971)
Rothemund, P.: Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Rothemund, P., Papadakis, N., Winfree, E.: Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biology 2(12), e424 (2004), pp. 2041–2053
Rothemund, P., Winfree, E.: The program-size complexity of self-assembled squares. In: Symposium on Theory of Computing, pp. 459–468 (2000)
Schulman, R., Winfree, E.: Programmable control of nucleation for algorithmic self-assembly. SIAM J. Comput. 39(4), 1581–1616 (2009)
Wang, H.: Proving theorems by pattern recognition II. Bell Syst. Tech. J. 40, 1–41 (1961)
Winfree, E.: On the computational power of DNA annealing and ligation. In: DNA Based Computers. DIMACS, vol. 27, pp. 199–221. Am. Math. Soc., Providence (1995)
Winfree, E.: Algorithmic self-assembly of DNA. Ph.D. thesis, California Institute of Technology (1998)
Winfree, E., Bekbolatov, R.: Proofreading tile sets: error correction for algorithmic self-assembly. In: Chen, J., Reif, J. (eds.) DNA Computing, Lecture Notes in Computer Science, vol. 2943, pp. 1980–1981. Springer, Berlin (2004)
Winfree, E., Liu, F., Wenzler, L., Seeman, N.: Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998)
Winfree, E., Yang, X., Seeman, N.: Universal computation via self-assembly of DNA: some theory and experiments. In: DNA Based Computers II. DIMACS, vol. 44, pp. 191–213. Am. Math. Soc., Providence (1996)
Yan, H., Feng, L., LaBean, T., Reif, J.: Parallel molecular computation of pair-wise XOR using DNA string tile. J. Am. Chem. Soc. 125(47), 14246–14247 (2003)
Yin, P., Choi, H., Calvert, C., Pierce, N.: Programming biomolecular self-assembly pathways. Nature 451(7176), 318–322 (2008)
Yin, P., Yan, H., Daniell, X., Turberfield, A., Reif, J.: A unidirectional DNA Walker moving autonomously along a linear track. Angew. Chem., Int. Ed. 116(37), 5014–5019 (2004)
Zhang, D., Turberfield, A., Yurke, B., Winfree, E.: Engineering entropy-driven reactions and networks catalyzed by DNA. Science 318, 1121–1125 (2007)
Acknowledgements
We thank our reviewers for pointing out errors and for comments that improve the readability of the paper. This work was supported by NSF EMT Grants CCF-0829797 and CCF-0829798.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chandran, H., Gopalkrishnan, N. & Reif, J. Tile Complexity of Approximate Squares. Algorithmica 66, 1–17 (2013). https://doi.org/10.1007/s00453-012-9620-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00453-012-9620-z