Abstract
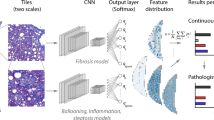
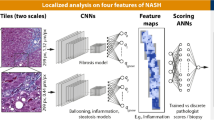
Non-alcoholic fatty liver disease (NAFLD) is one of the most frequent chronic liver diseases worldwide. Non-alcoholic steatohepatitis (NASH) is a progressive type of NAFLD that may cause cirrhosis, hepatocellular carcinoma, or almost mortality. Therefore, early diagnosis of the NASH is crucial. NASH is scored using the main histopathological features: ballooning, inflammation, steatosis, and fibrosis. The diagnosis of NASH by pathologists is time-consuming and can vary subjectively. On the other hand, several studies have reported deep learning approaches to enable fully automated NASH scoring. However, these studies suffer from limited labeled and imbalanced datasets. The purpose of this study to achieve fully automated NASH scoring with deep learning models that overcome data limitations. This study proposes an unsupervised transfer learning model for NASH scoring and fibrosis staging on a small-size dataset within two steps. In the first step, Convolutional Auto Encoder (CAE) is utilized as a deep feature extractor in an unsupervised manner during reconstruction. The second step is fine-tuning for classification consisting of the CAE encoder set as initial layers combined with fully connected layers and softmax. The proposed unsupervised transfer learning model is evaluated on a public NASH dataset. We compare the performance of the proposed network (CAE+classifier) with transfer learning models including Inception-v3, VGG16, ResNet-50. The proposed model has 94.87% AUC for ballooning, 89.47% AUC for inflammation, 96.15% AUC for steatosis, and 93.18% AUC for fibrosis. The proposed model is superior to transfer learning models (Inception-v3, VGG16, ResNet-50) with less parameter size and low computational complexity on a small NASH dataset.





Similar content being viewed by others
Data availibility
The datasets analysed during the current study are available in the https://osf.io/p48rd/ published by [8]. The data and code of the study are available from the corresponding author upon reasonable request.
References
Younossi ZM, Koenig AB, Abdelatif D et al (2016) Global epidemiology of nonalcoholic fatty liver disease-meta-analytic assessment of prevalence, incidence, and outcomes. Hepatology 64(1):73–84
Ge X, Zheng L, Wang M et al (2020) Prevalence trends in non-alcoholic fatty liver disease at the global, regional and national levels, 1990–2017: a population-based observational study. BMJ Open 10(8):e036663
Mitra S, De A, Chowdhury A (2020) Epidemiology of non-alcoholic and alcoholic fatty liver diseases. Transl Gastroenterol Hepatol 5
Sherif ZA, Saeed A, Ghavimi S et al (2016) Global epidemiology of nonalcoholic fatty liver disease and perspectives on us minority populations. Dig Dis Sci 61(5):1214–1225
Sugimoto K, Takei K (2015) Alcoholic liver disease and non-alcoholic fatty liver disease. J Jap Soc Gastroenterol 112(9):1641–1650. https://doi.org/10.11405/nisshoshi.112.1641
Takahashi Y, Fukusato T (2014) Histopathology of nonalcoholic fatty liver disease/nonalcoholic steatohepatitis. World J Gastroenterol: WJG 20(42):15539
Kleiner DE, Brunt EM, Van Natta M et al (2005) Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41(6):1313–1321
Heinemann F, Birk G, Stierstorfer B (2019) Deep learning enables pathologist-like scoring of nash models. Sci Rep 9(1):1–10
Pournik O, Alavian SM, Ghalichi L, et al (2014) Inter-observer and intra-observer agreement in pathological evaluation of non-alcoholic fatty liver disease suspected liver biopsies. Hepat Mon 14(1)
Taylor-Weiner A, Pokkalla H, Han L et al (2021) A machine learning approach enables quantitative measurement of liver histology and disease monitoring in nash. Hepatology 74(1):133–147
Becker H, Nettleton W, Meyers P et al (1964) Digital computer determination of a medical diagnostic index directly from chest x-ray images. IEEE Trans Biomed Eng 3:67–72
Lodwick GS, Keats TE, Dorst JP (1963) The coding of roentgen images for computer analysis as applied to lung cancer. Radiology 81(2):185–200
Chan HP, Lo SCB, Sahiner B et al (1995) Computer-aided detection of mammographic microcalcifications: pattern recognition with an artificial neural network. Med Phys 22(10):1555–1567
Lo SC, Lou SL, Lin JS et al (1995) Artificial convolution neural network techniques and applications for lung nodule detection. IEEE Trans Med Imaging 14(4):711–718
Sahiner B, Chan HP, Petrick N et al (1996) Classification of mass and normal breast tissue: a convolution neural network classifier with spatial domain and texture images. IEEE Trans Med Imaging 15(5):598–610
Wu Y, Giger ML, Doi K (1993) Artificial neural networks in mammography: application to decision making in the diagnosis of breast cancer. Radiology 187(1):81–87
Zhang W, Doi K, Giger ML (1996) An improved shift-invariant artificial neural network for computerized detection of clustered microcalcifications in digital mammograms. Med Phys 23(4):595–601
Pacal I, Karaboga D, Basturk A et al (2020) A comprehensive review of deep learning in colon cancer. Comput Biol Med 126:104003
Pacal I, Karaboga D (2021) A robust real-time deep learning based automatic polyp detection system. Comput Biol Med 134(104):519
Pacal I, Karaman A, Karaboga D et al (2022) An efficient real-time colonic polyp detection with yolo algorithms trained by using negative samples and large datasets. Comput Biol Med 141(105):031
Alici-Karaca D, Akay B, Yay A et al (2022) A new lightweight convolutional neural network for radiation-induced liver disease classification. Biomed Signal Process Control 73(103):463
Koohababni NA, Jahanifar M, Gooya A et al (2018) Nuclei detection using mixture density networks. International workshop on machine learning in medical imaging. Springer, New York, pp 241–248
Sirinukunwattana K, Raza SEA, Tsang YW et al (2016) Locality sensitive deep learning for detection and classification of nuclei in routine colon cancer histology images. IEEE Trans Med Imaging 35(5):1196–1206
Song TH, Sanchez V, Daly HE et al (2018) Simultaneous cell detection and classification in bone marrow histology images. IEEE J Biomed Health Inform 23(4):1469–1476
Shaban M, Awan R, Fraz MM et al (2020) Context-aware convolutional neural network for grading of colorectal cancer histology images. IEEE Trans Med Imaging 39(7):2395–2405
Reid D, Ternes K, Winowiecki L et al (2020) Germicidal irradiation of portable medical equipment: mitigating microbes and improving the margin of safety using a novel, point of care, germicidal disinfection pod. Am J Infect Control 48(1):103–105
Tang X, Gu X, Wang J et al (2020) A bearing fault diagnosis method based on feature selection feedback network and improved ds evidence fusion. IEEE Access 8:20523–20536
Jiao J, Zhao M, Lin J et al (2018) A multivariate encoder information based convolutional neural network for intelligent fault diagnosis of planetary gearboxes. Knowl-Based Syst 160:237–250
Han T, Liu C, Wu L et al (2019) An adaptive spatiotemporal feature learning approach for fault diagnosis in complex systems. Mech Syst Signal Process 117:170–187
Wen L, Li X, Gao L et al (2017) A new convolutional neural network-based data-driven fault diagnosis method. IEEE Trans Industr Electron 65(7):5990–5998
Wang S, Xiang J, Zhong Y et al (2018) Convolutional neural network-based hidden markov models for rolling element bearing fault identification. Knowl-Based Syst 144:65–76
Jing L, Zhao M, Li P et al (2017) A convolutional neural network based feature learning and fault diagnosis method for the condition monitoring of gearbox. Measurement 111:1–10
Verstraete D, Ferrada A, Droguett EL, et al (2017) Deep learning enabled fault diagnosis using time-frequency image analysis of rolling element bearings. Shock Vib 2017
Han T, Liu C, Yang W et al (2019) A novel adversarial learning framework in deep convolutional neural network for intelligent diagnosis of mechanical faults. Knowl-Based Syst 165:474–487
Islam MM, Kim JM (2019) Automated bearing fault diagnosis scheme using 2d representation of wavelet packet transform and deep convolutional neural network. Comput Ind 106:142–153
Stojanovic V, Prsic D (2020) Robust identification for fault detection in the presence of non-gaussian noises: application to hydraulic servo drives. Nonlinear Dyn 100(3):2299–2313
Dong X, He S, Stojanovic V (2020) Robust fault detection filter design for a class of discrete-time conic-type non-linear markov jump systems with jump fault signals. IET Control Theory Appl 14(14):1912–1919
Stojanovic V, He S, Zhang B (2020) State and parameter joint estimation of linear stochastic systems in presence of faults and non-gaussian noises. Int J Robust Nonlinear Control 30(16):6683–6700
Tao H, Wang P, Chen Y et al (2020) An unsupervised fault diagnosis method for rolling bearing using stft and generative neural networks. J Franklin Inst 357(11):7286–7307
Yan R, Ren F, Wang Z et al (2020) Breast cancer histopathological image classification using a hybrid deep neural network. Methods 173:52–60
Man R, Yang P, Xu B (2020) Classification of breast cancer histopathological images using discriminative patches screened by generative adversarial networks. IEEE Access 8:155362–155377
Xue Y, Ye J, Zhou Q et al (2021) Selective synthetic augmentation with histogan for improved histopathology image classification. Med Image Anal 67(101):816
Ahmad N, Asghar S, Gillani SA (2022) Transfer learning-assisted multi-resolution breast cancer histopathological images classification. Vis Comput 38(8):2751–2770
Shao Z, Bian H, Chen Y et al (2021) Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv Neural Inf Process Syst 34:2136–2147
Abdelsamea MM, Zidan U, Senousy Z et al (2022) A survey on artificial intelligence in histopathology image analysis. Wiley Interdiscip Rev Data Min Knowl Discov 12:e1474
Kaur S, Singla J, Nkenyereye L et al (2020) Medical diagnostic systems using artificial intelligence (ai) algorithms: principles and perspectives. IEEE Access 8:228049–228069
Altaf F, Islam SM, Akhtar N et al (2019) Going deep in medical image analysis: concepts, methods, challenges, and future directions. IEEE Access 7:99540–99572
Deng J, Dong W, Socher R, et al (2009) Imagenet: a large-scale hierarchical image database. In: 2009 IEEE conference on computer vision and pattern recognition. IEEE, pp 248–255
Szegedy C, Vanhoucke V, Ioffe S, et al (2016) Rethinking the inception architecture for computer vision. In: Proceedings of the IEEE conference on computer vision and pattern recognition. pp 2818–2826
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556
He K, Zhang X, Ren S, et al (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition. pp 770–778
Yu Y, Wang J, Ng CW et al (2018) Deep learning enables automated scoring of liver fibrosis stages. Sci Rep 8(1):1–10
Arjmand A, Angelis CT, Christou V et al (2020) Training of deep convolutional neural networks to identify critical liver alterations in histopathology image samples. Appl Sci 10(1):42
Roy M, Wang F, Vo H et al (2020) Deep-learning-based accurate hepatic steatosis quantification for histological assessment of liver biopsies. Lab Invest 100(10):1367–1383
Fu X, Liu T, Xiong Z et al (2018) Segmentation of histological images and fibrosis identification with a convolutional neural network. Comput Biol Med 98:147–158
Sun L, Marsh JN, Matlock MK et al (2020) Deep learning quantification of percent steatosis in donor liver biopsy frozen sections. EBioMedicine 60(103):029
Levy JJ, Salas LA, Christensen BC, et al (2019) Pathflowai: a high-throughput workflow for preprocessing, deep learning and interpretation in digital pathology. In: Pacific symposium on biocomputing 2020. World Scientific. pp 403–414
Qu H, Minacapelli CD, Tait C et al (2021) Training of computational algorithms to predict NAFLD activity score and fibrosis stage from liver histopathology slides. Comput Methods Programs Biomed 207(106):153
Rumelhart DE, Hinton GE, Williams RJ (1985) Learning internal representations by error propagation. California Univ San Diego La Jolla Inst for Cognitive Science, Tech. rep
Hinton GE, Salakhutdinov RR (2006) Reducing the dimensionality of data with neural networks. Science 313(5786):504–507
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521(7553):436–444
Bloice MD, Roth PM, Holzinger A (2019) Biomedical image augmentation using Augmentor. Bioinformatics 35(21):4522–4524. https://doi.org/10.1093/bioinformatics/btz259
Chollet F, et al (2015) Keras. https://github.com/fchollet/keras
Paszke A, Gross S, Massa F, et al (2019) Pytorch: An imperative style, high-performance deep learning library. In: Advances in neural information processing systems 32. Curran Associates, Inc. p 8024–8035, http://papers.neurips.cc/paper/9015-pytorch-an-imperative-style-high-performance-deep-learning-library.pdf
Al-Kababji A, Bensaali F, Dakua SP (2022) Scheduling techniques for liver segmentation: Reducelronplateau vs onecyclelr. arXiv preprint arXiv:2202.06373
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no potential conflicts of interest regarding any financial support, research, authorship and publication of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Karagoz, M.A., Akay, B., Basturk, A. et al. An unsupervised transfer learning model based on convolutional auto encoder for non-alcoholic steatohepatitis activity scoring and fibrosis staging of liver histopathological images. Neural Comput & Applic 35, 10605–10619 (2023). https://doi.org/10.1007/s00521-023-08252-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-023-08252-2