Abstract
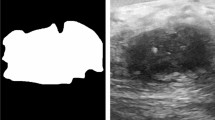
Automated Breast Ultrasound is a highly advanced breast tumor detection modality that produces hundreds of 2D slices in each scan. However, this large number of slices poses a significant burden for physicians to review. This paper proposes a novel 2.5D tumor detection model, named “ABUSDet,” to assist physicians in automatically reviewing ABUS images and predicting the locations of breast tumors in images. At the core of this approach, a sequence of data blocks partitioned from a pre-processed 3D volume are fed to a 2.5D tumor detection model, which outputs a sequence of 2D tumor candidates. An aggregation module then clusters the 2D tumor candidates to produce the ultimate 3D coordinates of the tumors. To further improve the accuracy of the model, a novel mechanism for training deep learning models, called “Deliberate Training,” is proposed. The proposed model is trained and tested on a dataset of 87 patients with 235 ABUS volumes. It achieves sensitivities of 77.94%, 75.49%, and 65.19% at FPs/volume of 3, 2, and 1, respectively. Compared with the 2D and 3D object detection models, the proposed ABUSDet model achieves the highest sensitivity with relatively low false-positive rates.
Graphical abstract














Similar content being viewed by others
Data availability
Due to ethical concerns, supporting data cannot be made openly available.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin 71(3):209–249
Mridha MF, Hamid MA, Monowar MM, Keya AJ, Ohi AQ, Islam MR, Kim J-M (2021) A comprehensive survey on deep-learning-based breast cancer diagnosis. Cancers 13(23):6116
Duggan C, Trapani D, Ilbawi AM, Fidarova E, Laversanne M, Curigliano G, Bray F, Anderson BO (2021) National health system char acteristics, breast cancer stage at diagnosis, and breast cancer mortality: a population-based analysis. Lancet Oncol 22(11):1632–1642
Zamora K, Allen E, Hermecz B (2021) Contrast mammography in clinical practice: Current uses and potential diagnostic dilemmas. Clin Imaging 71:126–135
Zhang Z, Wang W, Wang X, Yu X, Zhu Y, Zhan H, Chen Z, Li B, Huang J (2020) Breast-specific gamma imaging or ultrasonography as adjunct imaging diagnostics in women with mammographically dense breasts. Eur Radiol 30(11):6062–6071
Luczyńska E, Pawlak M, Popiela T, Rudnicki W (2022) The role of abus in the diagnosis of breast cancer. J Ultrasonogr 22(89):76–85
Moon WK, Shen Y-W, Bae MS, Huang C-S, Chen J-H, Chang R-F (2013) Computer-aided tumor detection based on multi-scale blob detection algorithm in automated breast ultrasound images. IEEE Trans Med Imaging 32(7):1191–1200. https://doi.org/10.1109/TMI.2012.2230403
Tan T, Platel B, Mus R, Tabár L, Mann RM, Karssemeijer N (2013) Computer-aided detection of cancer in automated 3-d breast ultrasound. IEEE Transactions on Medical Imaging 32(9):1698–1706. https://doi.org/10.1109/TMI.2013.2263389
Lo C-M, Chen R-T, Chang Y-C, Yang Y-W, Hung M-J, Huang C-S, Chang R-F (2014) Multi-dimensional tumor detection in automated whole breast ultrasound using topographic watershed. IEEE Trans Med Imaging 33(7):1503–1511. https://doi.org/10.1109/TMI.2014.2315206
Chiang T-C, Huang Y-S, Chen R-T, Huang C-S, Chang R-F (2019) Tumor detection in automated breast ultrasound using 3-d cnn and prioritized candidate aggregation. IEEE Trans Med Imaging 38(1):240–249. https://doi.org/10.1109/TMI.2018.2860257
Wang Y, Wang N, Xu M, Yu J, Qin C, Luo X, Yang X, Wang T, Li A, Ni D (2020) Deeply-supervised networks with threshold loss for cancer detection in automated breast ultrasound. IEEE Trans Med Imaging 39(4):866–876. https://doi.org/10.1109/TMI.2019.2936500
Zhou Y, Chen H, Li Y, Wang S, Cheng L, Li J (2021) 3d multi-view tumor detection in automated whole breast ultrasound using deep convolutional neural network. Expert Syst Applic 168:114410. https://doi.org/10.1016/j.eswa.2020.114410
Li Y, Wu W, Chen H, Cheng L, Wang S (2020) 3d tumor detection in automated breast ultrasound using deep convolutional neural network. Med Phys 47(11):5669–5680
Xiang H, Huang Y-S, Lee C-H, Chien T-YC, Lee C-K, Liu L, Li A, Lin X, Chang R-F (2021) 3-d res-capsnet convolutional neural network on automated breast ultrasound tumor diagnosis. Eur J Radiol 138:109608
Zhang Z, Li Y, Wu W, Chen H, Cheng L, Wang S (2021) Tumor detection using deep learning method in automated breast ultrasound. Biomed Signal Proc Control 68:102677. https://doi.org/10.1016/j.bspc.2021.102677
Hu J, Shen L, Albanie S, Sun G, Wu E (2020) Squeeze-and-excitation networks. IEEE Trans Pattern Anal Mach Intell 42(8):2011–2023
Ester M, Kriegel H-P, Sander J, Xu X et al (1996) A density-based algorithm for discovering clusters in large spatial databases with noise. Kdd 96:226–231
Ayana G, Dese K, Choe S-W (2021) Transfer learning in breast cancer diagnoses via ultrasound imaging. Cancers 13(4):738
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition, IEEE, pp. 770–778
Deng J, Dong W, Socher R, Li L-J, Li K, Fei-Fei L (2009) Imagenet: A large-scale hierarchical image database. In: 2009 IEEE Conference on Computer Vision and Pattern Recognition, pp. 248–255. https://doi.org/10.1109/CVPR.2009.5206848
Anders Ericsson K (2008) Deliberate practice and acquisition of expert performance: a general overview. Acad Emerg Med 15(11):988–994
Neubeck A, Van Gool L (2006) Efficient non-maximum suppression. In: 18th International Conference on Pattern Recognition (ICPR’06), IEEE Computer Society, vol. 3, pp. 850–855
Felzenszwalb PF, Girshick RB, McAllester D, Ramanan D (2010) Object detection with discriminatively trained part-based models. IEEE Trans Pattern Anal Mach Intell 32(9):1627–1645
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Killeen T, Lin Z, Gimelshein N, Antiga L et al (2019) Pytorch: An imperative style, high-performance deep learning library. Adv Neural Inform Proc Syst 8026–8037
Chakraborty DP (1989) Maximum likelihood analysis of free-response receiver operating characteristic (froc) data. Med Phys 16(4):561–568
Lobo JM, Jiḿenez-Valverde A, Real R (2008) Auc: a misleading measure of the performance of predictive distribution models. Global Ecol Biogeogr 17(2):145–151
Acknowledgments
We want to thank the Traditional Chinese Medicine Hospital of Guangdong Province for providing data for this study.
Funding
This work is partially supported by the National Natural Science Foundation of China (61976037), the Yongjiang Technology Innovation Project (2022A-097-G), and the Ningbo 2025 Key R&D Project (2023Z223).
Author information
Authors and Affiliations
Contributions
Xudong Song: Investigation, data curation, writing the manuscript, and conducting the experiments. Xiaoyang Lu: Replicated Ref [8]. Gengfa Fang: Investigation, data curation, project administration, supervision, writing, review, commentary, and revision. Xiangjian He: Funding acquisition, supervision, writing, review, commentary, and revision. Xiaochen Fan: Data analysis, review, commentary. Le cai: Data analysis, review, commentary. Wenjing Jia: Writing, review, commentary, and revision. Zumin Wang: Writing, review, commentary, and revision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, X., Lu, X., Fang, G. et al. ABUSDet: A Novel 2.5D deep learning model for automated breast ultrasound tumor detection. Appl Intell 53, 26255–26269 (2023). https://doi.org/10.1007/s10489-023-04785-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10489-023-04785-0