Abstract
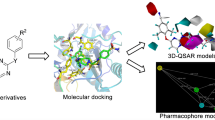
To find useful information for discovering dual functional inhibitors against both wild type (WT) and K103N mutant reverse transcriptases (RTs) of HIV-1, molecular docking and 3D-QSAR approaches were applied to a set of twenty-five 4,1-benzoxazepinone analogues of efavirenz (SUSTIVA®), some of them are active against the two RTs. 3D-QSAR models were constructed, based on their binding conformations determined by molecular docking, with r 2 cv values ranging from 0.656 to 0.834 for CoMFA and CoMSIA, respectively. The models were then validated to be highly predictive and extrapolative by inhibitors in two test sets with different molecular skeletons. Furthermore, CoMFA models were found to be well matched with the binding sites of both WT and K103N RTs. Finally, a reasonable pharmacophore model of 4,1-benzoxazepinones were established. The application of the model not only successfully differentiated the experimentally determined inhibitors from non-inhibitors, but also discovered two potent inhibitors from the compound database SPECS. On the basis of both the 3D-QSAR and pharmacophore models, new clues for discovering and designing potent dual functional drug leads against HIV-1 were proposed: (i) adopting positively charged aliphatic group at the cis-substituent of C3; (ii) reducing the electronic density at the position of O4; (iii) positioning a small branched aliphatic group at position of C5; (iv) using the negatively charged bulky substituents at position of C7.
Similar content being viewed by others
References
Brian GT, Summers MF (1999) J Mol Biol 285:1
Emerman M, Malim MH (1998) Science 280:1880
Mitsuya H, Broder S (1987) Nature 325:773
Esnouf R, Ren J, Ross C, Jones Y, Stammers D, Stuart D (1995) Nat Struct Biol 2:303
Jones PS (1998) Antiviral Chem Chemother 9:283
Pedersen OS, Pedersen EB (1999) Antiviral Chem Chemother 10:285
Vella S, Palmisano L (2000) Antiviral Res 45:1
Bardsley-Elliot A, Perry CM (2000) Pediatric Drugs 2:373
Corbett JW, Ko SS, Rodgers JD, Gearhart LA, Magnus NA, Bacheler LT, Diamond S, Jeffrey S, Klabe RM, Cordova BC, Garber S, Logue K, Trainor GL, Anderson PS, Erickson-Viitanen SK (2000) J Med Chem 43:2019
Moyle G (2001) Drugs 61:19
Lee K, Gulick RM (2001) Curr Infec Dis Rep 3:193
Adkins JC, Nobel S (1998) Drugs 56:1055
Robert WB Jr (2001) Expert Opin Investig Drugs 10:1423
Young SD, Britcher SF, Tran LO, Payne LS, Lumma WC, Lyle TA, Huff JR, Anderson PS, Olsen DB, Carroll SS (1995) Antimicrob Agents Chemother 39:2602
Lindberg J, Sigurdsson S, Lowgren S, Andersson HO, Sahlberg C, Noreen R, Fridborg K, Zhang H, Unge T (2002) Eur J Biochem 269:1670
Ren J, Nichols C, Bird L, Chamberlain P, Weaver K, Short S, Stuart DI, Stammers DK (2001) J Mol Biol 312:795
Jay AM, David DC, Abdul M, Beverly CC, Ronald MK, Steven PS (2001) Bioorg Med Chem Lett 11:619
Chamberlain PP, Ren J, Nichols CE, Douglas L, Lennerstrand J, Larder BA, Stuart DI, Stammers DK (2002) J Virol 76:10015
Medina-Franco JL, Rodríguez-Morales S, Juárez-Gordiano C, Hernández-Campos A, Castillo R (2004) J Comput Aided Mol Des 18:345
De Clercq E (2001) Curr Med Chem 8:1543
Ding J, Das K, Moereels H, Koymans L, Andries K, Paul AJ, Atephen J, Huges H, Arnold E (1995) Struct Biol 2:407
Ren J, Milton J, Weaver KL, Short SA, Stuart DI, Stammers DK (2000) Structure 8:1089
Kohlstaedt LA, Wang J, Friedman JM, Rice PA, Steitz TA (1992) Science 256:1783
Ding J, Das K, Tantillo C, Zhang W, Clark ADJ, Jessen S, Lu X, Hsiou Y, Jacobo-Molina A, Andries K (1995) Structure 3:365
Mao C, Sudbeck EA, Venkatachalam TK, Uckun FM (1999b) Antiviral Chem Chemother 10:233
Cocuzza AJ, Chidester DR, Cordova BC, Klabe RM, Jeffrey S, Diamond S, Weigelt CA, Ko SS, Bacheler LT, Erickson-Viitanen SK, Rodgers JD (2001) Bioorg Med Chem Lett 11:1389
Cramer RD, Patterson DE, Bunce JD (1988) J Am Chem Soc 110:5959
Klebe G, Abraham U (1999) J Comput Aided Mol Des 13:1
Liu G, Zhang Z, Luo X, Shen J, Liu H, Shen X, Chen K, Jiang H (2004) Bioorg Med Chem 12:4147
Powell MJD (1977) Math Programming 12:241
Marsili M, Gasteiger J (1980) Croat Chem Acta 53:601
Gasteiger J, Marsili M (1980) Tetrahedron 36:3219
Morris GM, Goodsell DS, Huey R, Hart WE, Halliday S, Belew R, Olson AJ (1999) AutoDock, Version 3.0.3. The Scripps Research Institute, Molecular Graphics Laboratory, Department of Molecular Biology
Wallace AC, Laskowski RA, Thornton JM (1995) Protein Eng 8:127
Kim KS, Tarakeshwar P, Lee JY (2000) Chem Rev 100:4145
Pungpo P, Hannongbua S (2000) J Mol Graphics Mod 18:581
Corbett JW, Ko SS, Rodgers JD, Gearhart LA, Magnus NA, Bacheler LT, Diamond S, Jeffrey S, Klabe RM, Cordova BC, Garber S, Logue K, Trainor GL, Anderson PS, Erickson-viitanen SK (2000) J Med Chem 43:2019
Campiani G, Ramunno A, Maga G, Nacci V, Fattorusso C, Novellino E (2002) Curr Pharma Des 8:615
Urabe T, Sano K, Tanno M, Mizoguchi J, Otani M, Lee MH, Takasaki T, Kusakabe H, Imagawa DT, Nakai M (1992) J Virol Methods 40:145
Acknowledgment
The authors acknowledge the financial supports Shanghai Key Basic R&D Program (grants 03DZ19228 and 05JC14092), and 863 program (grant 2003AA235010).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, Z., Zheng, M., Du, L. et al. Towards discovering dual functional inhibitors against both wild type and K103N mutant HIV-1 reverse transcriptases: molecular docking and QSAR studies on 4,1-benzoxazepinone analogues. J Comput Aided Mol Des 20, 281–293 (2006). https://doi.org/10.1007/s10822-006-9050-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-006-9050-6