Abstract
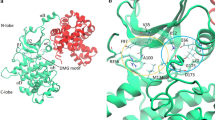
Interests in CDK2 and CDK5 have stemmed mainly from their association with cancer and neuronal migration or differentiation related diseases and the need to design selective inhibitors for these kinases. Molecular dynamics (MD) simulations have not only become a viable approach to drug design because of advances in computer technology but are increasingly an integral part of drug discovery processes. It is common in MD simulations of inhibitor/CDK complexes to exclude the activator of the CDKs in the structural models to keep computational time tractable. In this paper, we present simulation results of CDK2 and CDK5 with roscovitine using models with and without their activators (cyclinA and p25). While p25 was found to induce slight changes in CDK5, the calculations support that cyclinA leads to significant conformational changes near the active site of CDK2. This suggests that detailed and structure-based inhibitor design targeted at these CDKs should employ activator-included models of the kinases. Comparisons between P/CDK2/cyclinA/roscovitine and CDK5/p25/roscovitine complexes reveal differences in the conformations of the glutamine around the active sites, which may be exploited to find highly selective inhibitors with respect to CDK2 and CDK5.








Similar content being viewed by others
Abbreviations
- (CDK):
-
Cyclin-Dependent Kinase
- P/CDK2/cyclinA/roscovitine:
-
Thr160−phosphated/CDK2/cyclinA/roscovitine
References
Harper JW, Adams PD (2001) Chem Rev 101:2511
Garrett MD, Fattaey A (1999) Curr Opin Genet Dev 9:104
Webster KR (1998) Expert Opin Investig Drugs 7:865
Nguyen MD, Julien JP (2003) Neurosignals 12:215
Lau LF, Ahlijanian MK (2003) Neurosignals 12:209
Smith PD, Crocker SJ, Jackson-Lewis V, Jordan-Sciutto KL, Hayley S (2003) Proc Natl Acad Sci USA 100:13650
Bu B, Li J, Davies P, Vincent I (2002) J Neurosci 22:6515
Wang J, Liu S, Fu Y, Wang JH, Lu Y (2003) Nat Neurosci 6:1039
Hunter T, Pines J (1994) Cell 79:573
Sherr C (1996) Science 274:1672
Norbury C, Nurse P (1992) Annu Rev Biochem 61:441
De Azevedo WF Jr, Mueller-Dieckmann HJ, Schulze-Gahmen U, Worland PJ, Sausville E, Kim SH (1996) Proc Natl Acad Sci USA 93:2735
De Azevedo WF Jr, Leclerc S, Meijer L, Havlicek L, Strnad M, Kim SH (1997) Eur J Biochem 243:518
Jeffrey PD, Russo AA, Polyak K, Gibbs E, Hurwitz J, Massague J, Pavletich NP (1995) Nature 376:313
Patrick GN, Zhou P, Kwon YT, Howley PM, Tsai L-H (1998) J Biol Chem 273:24057
Patrick GN, Zukerberg L, Nikolic M, de la Monte S, Dikkes P, Tsai L-H (1999) Nature 402:615
Tarricone C, Dhavan R, Peng J, Areces LB, Tsai L-H, Musacchio A (2001) Mol Cell 8:657
Russo A, Jeffrey PD, Pavletich NP (1996) Nat Struct Biol 3:696
Morgan DO (1996) Curr Opin Cell Biol 8:767
Cavalli A, Dezi C, Folkers G, Scapozza L, Recanatini M (2001) Proteins Struct Funct Genet 45:478
Otyepka M, Kříž Z, Koča J (2002) J Biomol Struct Dyn 20:141
De Azevedo WF Jr, Gaspar RT, Canduri F, Camera JC Jr, Silveira NJF (2002) Biochem Biophys Res Commun 297:1154
Cavalli A, Vivo MD, Recanatini M (2003) Chem Commum 1308
Sims PA, Wong CF, McCammon JA (2003) J Med Chem 46:3314
Park H, Yeom MS, Lee S (2004) ChemBioChem 5:1662
Bártová I, Otyepka M, Kříž Z, Koča (2004) Protein Sci 13:1449
Bártová I, Otyepka M, Kříž Z, Koča J (2005) Protein Sci 14:445
Jiang Y-J, Zou J-W, Gui C-S (2005) J Mol Model 11:509
García-Sosa AT, Mancera RL (2006) J Mol Model 12:422
Davies TG, Tunnah P, Meijer L, Marko D, Eisenbrand G, Endicott JA, Noble MEM (2001) Structure 9:389
Leclerc S, Garnier M, Hoessel R, Marko D, Bibb JA, Snyder GL, Greengard P, Biernati J, Wui Y-Z, Mandelkowi E-M, Eisenbrand G, Meijer L (2001) J Biol Chem 276:251
Sausville EA (2002) Trends Mol Med 8:S32
Noble MEM, Endicott JA, Johnson LN (2004) Science 303:1800
Noble M, Barrett P, Endicott J, Johnson L, McDonnell J, Robertson G, Zawaira A (2005) Biochimica et Biophysica Acta 1754:58
Huew A, Mazitschek R, Giannis A (2003) Angew Chem Int Ed 42:2122
Sielecki M, Boylan JF, Benfield PA, Trainor GL (2000) J Med Chem 43:1
Knockaert P, Greengard L, Meijer L (2002) Trends Pharmacol Sci 23:417
Lau LF, Seymour PA, Sanner MA, Schachter JB (2002) J Mol Neurosci 19:267
Crews CM, Mohan R (2000) Curr Opin Chem Biol 4:47
Meijer L, Borgne A, Mulner O, Chong JP, Blow JJ, Inagaki N (1997) Eur J Biochem 243:527
Ljungman M, Paulsen MT (2001) Mol Pharmacol 60:785
Bach S, Knockaert M, Reinhardt J, Lozach O, Schmitt S, Baratte B, Koken M, Coburn SP, Tang L, Jiang T, Liang DC, Galons H, Dierick JF, Pinna LA, Meggio F, Totzke F, Schachtele C, Lerman AS, Carnero A, Wan Y, Gray N, Meijer L (2005) J Biol Chem 280:31208
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM Jr, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) J Am Chem Soc 117:5179
Besler BH, Merz KM, Kollman PA (1990) J Comput Chem 11:431
Fox T, Kollman PA (1998) J Phys Chem B 102:8070
Case DA, Pearlman DA, Caldwell JW, Cheatham TE III, Wang J, Ross WS, Simmerling CL, Darden TA, Merz KM, Stanton RV, Cheng AL, Vincent JJ, rowley M, Tsui V, Gohlke H, Radme RJ, Duan Y, Pitera J, Massova I, Seibel GL, Singh UC, Weiner PK, Kollman PA (2002) AMBER 7. University of California, San Francisco
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) J Chem Phys 79:926
Darden T, York D, Pedersen L (1993) J Chem Phys 98:10089
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) J Comput Phys 23:327
Morris GM, Goodsell DS, Huey R, Hart WE, Halliday S, Belew R, Olson AJ (1999) Autodock Version 3.0.3. Molecular Graphics Laboratory, Department of Molecular Biology, The Scripps Research Institute
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) J Comput Chem 19:1639
Massova I, Kollman PA (2000) Perspect Drug Discov 18:113
Connolly ML (1983) J Appl Cryst 16:548
Sitkoff D, Sharp KA, Honig B (1994) J Phys Chem 98:1978
Lemaitre V, Ali R, Kim C, Watts A, Fischer WB (2004) FEBS Lett 563:75
Acknowledgement
Prof. Meijer Laurent in France and Dr. Sung-Hou KIM in University of Berkeley are gratefully acknowledged for generously offering the crystal structures of the CDK2/R-roscovitine complex.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, B., Tan, V.B.C., Lim, K.M. et al. Molecular dynamics simulations on the inhibition of Cyclin-Dependent Kinases 2 and 5 in the presence of activators. J Comput Aided Mol Des 20, 395–404 (2006). https://doi.org/10.1007/s10822-006-9081-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-006-9081-z