Abstract
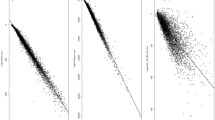
In protein structure prediction, a considerable number of models are usually produced by either the Template-Based Method (TBM) or the ab initio prediction. The purpose of this study is to find the critical parameter in assessing the quality of the predicted models. A non-redundant template library was developed and 138 target sequences were modeled. The target sequences were all distant from the proteins in the template library and were aligned with template library proteins on the basis of the transformation matrix. The quality of each model was first assessed with QMEAN and its six parameters, which are C_β interaction energy (C_beta), all-atom pairwise energy (PE), solvation energy (SE), torsion angle energy (TAE), secondary structure agreement (SSA), and solvent accessibility agreement (SAE). Finally, the alignment score (score) was also used to assess the quality of model. Hence, a total of eight parameters (i.e., QMEAN, C_beta, PE, SE, TAE, SSA, SAE, score) were independently used to assess the quality of each model. The results indicate that SSA is the best parameter to estimate the quality of the model.

Similar content being viewed by others
Abbreviations
- C_beta:
-
C_β interaction energy
- PE:
-
All-atom pairwise energy
- SE:
-
Solvation energy
- TAE:
-
Torsion angle energy
- SSA:
-
Secondary structure agreement
- SAA:
-
Solvent accessibility agreement
- TBM:
-
Template-based method
- CASP:
-
Critical assessment of protein structure prediction
References
Murzin AG (2001) Progress in protein structure prediction. Nat Struct Biol 8:110–112
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Sánchez R, Sali A (1997) Advances in comparative protein-structure modelling. Curr Opin Struct Biol 7:206–214
Bowie JU, Luthy R, Eisenberg D (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253:164–170
Panchenko AR, Marchler-Bauer A, Bryant SH (2000) Combination of threading potentials and sequence profiles improves fold recognition. J Mol Biol 296:1319–1331
Xu D, Zhang J, Roy A, Zhang Y (2011) Automated protein structure modeling in CASP9 by I-TASSER pipeline combined with QUARK-based ab initio folding and FG-MD-based structure refinement. Proteins: Struct Funct Bioinform 79:147–160
Pillardy J, Czaplewski C, Liwo A, Lee J, Ripoll DR, Kaźmierkiewicz R, Oldziej S, Wedemeyer WJ, Gibson KD, Arnautova YA, Saunders J, Ye YJ, Scheraga HA (2001) Recent improvements in prediction of protein structure by global optimization of a potential energy function. Proc Natl Acad Sci USA 98:2329–2333
Simons KT, Strauss C, Baker D (2001) Prospects for ab initio protein structural genomics. J Mol Biol 306:1191–1199
Kolinski A, Skolnick J (1998) Assembly of protein structure from sparse experimental data: an efficient Monte Carlo model. Proteins 32:475–494
Elofsson A (2002) A study on protein sequence alignment quality. Proteins 46:330–339
Pandit SB, Zhang Y, Skolnick J (2006) TASSER-Lite: an automated tool for protein comparative modeling. Biophys J 91:4180–4190
Kryshtafovych A, Fidelis K (2009) Protein structure prediction and model quality assess. Drug Discovery Today 14:386–393
Pearson WR, Sierk ML (2005) The limits of protein sequence comparison. Curr Opin Struct Biol 15:254–260
Levitt M, Gerstein M (1998) A unified statistical framework for sequence comparison and structure comparison. Proc Natl Acad Sci USA 95:5913–5920
Siew N, Elofsson A, Rychlewski L, Fischer D (2000) MaxSub: an automated measure for the assessment of protein structure prediction quality. Bioinformatics 16:776–785
Rohl CA, Strauss CEM, Misura KMS, Baker D (2004) Protein structure prediction using Rosetta. Methods Enzymol 383:66–93
Zhang Y, Skolnick J (2004) Automated structure prediction of weakly homologous proteins on a genomic scale. Proc Natl Acad Sci USA 101:7594–7599
Contreras-Moreira B, Fitzjohn PW, Bates PA (2003) In silico protein recombination:enhancing template and sequence alignment selection for comparative protein modelling. J Mol Biol 328:593–608
John B, Sali A (2003) Comparative protein structure modeling by iterative alignment, model building and model assessment. Nucleic Acids Res 31:3982–3992
Benkert P et al (2008) QMEAN: a comprehensive scoring function for model quality assessment. Proteins 71:261–277
Wallner B, Fang H, Elofsson A (2003) Automatic consensus-based fold recognition using Pcons, ProQ, and Pmodeller. Proteins 53:534–541
Li J, Fang H (2012) Substitution transformation of score matrix for improving alignment quality of local sequence of distantly related proteins. Current Bioinformtics 7:35–42
Murzin AG, Brenner SE, Hubbard T (1995) SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol 247:536–540
Fang H, Wallner B, Lundström J, von Wowern C, Elofsson A (2001) Improved fold recognition by using the Pcons consensus approach. Protein Struct Prediction: Bioinform Approach 1:397–416
Chen H, Kihara D (2008) Estimating quality of template-based protein models by alignment stability. Proteins 71:1255–1274
Rost B (2001) Review: protein secondary structure prediction continues to rise. J Struct Biol 134:204–218
Ortiz AR, Kolinski A, Rotkiewicz P, Ilkowski B, Skolnick J (1999) Ab initio folding of proteins using restraints derived from evolutionary information. Proteins 37:177–185
Eyrich VA, Standley DM, Felts AK, Friesner RA (1999) Protein tertiary structure prediction using a branch and bound algorithm. Proteins 35:41–57
Lomize AL, Pogozheva ID, Mosberg HI (1999) Prediction of protein structure: the problem of fold multiplicity. Proteins 37:199–203
Chen CC, Singh JP, Altman RB (1999) Using imperfect secondary structure predictions to improve molecular structure computations. Bioinformatics 15:53–65
Samudrala R, Xia Y, Huang E, Levitt M (1999) Ab initio protein structure prediction using a combined hierarchical approach. Proteins 37:194–198
Samudrala R, Huang ES, Koehl P, Levitt M (2000) Constructing side chains on near-native main chains for ab initio protein structure prediction. Protein Eng 13:453–457
Acknowledgments
This work was supported by Grants from the National Natural Science Foundation of China (No. 31171386 and 31371399), the Jiangsu Province Natural Science Foundation (No. BK2011093).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, J., Fang, H. A comparison of different functions for predicted protein model quality assessment. J Comput Aided Mol Des 30, 553–558 (2016). https://doi.org/10.1007/s10822-016-9924-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-9924-1