Abstract
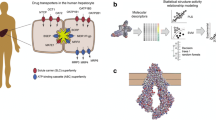
The parallel artificial membrane permeability assay (PAMPA), a non-cellular lab-based assay, is extensively used to measure the permeability of pharmaceutical compounds. PAMPA experiments provide a working mimic of a molecule passing through cells and PAMPA values are widely used to estimate drug absorption parameters. There is an increased interest in developing computational methods to predict PAMPA permeability values. We developed an in silico model to predict the permeability of compounds based on the PAMPA assay. We used the three-dimensional reference interaction site model (3D-RISM) theory with the Kovalenko–Hirata (KH) closure to calculate the excess chemical potentials of a large set of compounds and predicted their apparent permeability with good accuracy (mean absolute error or MAE = 0.69 units) when compared to a published experimental data set. Furthermore, our in silico PAMPA protocol performed very well in the binary prediction of 288 compounds as being permeable or impermeable (precision = 94%, accuracy = 93%). This suggests that our in silico protocol can mimic the PAMPA assay and could aid in the rapid discovery or screening of potentially therapeutic drug leads that can be delivered to a desired tissue.



Similar content being viewed by others
References
Kansy M, Senner F, Gubernator L (1998) J Med Chem 41:1007–1010
Kansy M, Fischer H, Kratzat K, Senner F, Wagner B, Parrilla I (2001) In: Testa B, van de Waterbeemd H, Folkers G, Guy R (eds) Pharmacokinetic optimization in drug research: biological, physicochemical, and computational strategies. Verlag Helvetica Chimica Acta, Zürich/Wiley-VCG, Weinheim
Di L, Kerns EH (2003) Curr Opin Chem Biol 7:402–408
Veber DF, Johnson SR, Cheng H-Y, Smith BR, Ward KW, Kopple KD (2002) J Med Chem 45:2615–2623
Kerns EH (2001) J Pharm Sci 90:1838–1858
Avdeef A, Bendels S, Di L, Faller B, Kansy M, Sugano K, Yamauchi Y (2007) J Pharm Sci 96:2893–2909
Bermejo M, Avdeef A, Ruiz A, Nalda R, Ruell JA, Tsinman O, González I, Fernández C, Sánchez G, Garrigues TM, Merino V (2004) Eur J Pharm Sci 21:429–441
Avdeef A, Artursson P, Neuhoff S, Lazorova L, Gråsjӧ J, Tavelin S (2005) Eur J Pharm Sci 24:333–349
Wohnsland F, Faller B (2001) J Med Chem 44:923–930
Sugano K, Hamada H, Machida M, Ushio H (2001) J Biomol Screen 6:189–196
Fujikawa M, Ano R, Nakao K, Shimizu R, Akamatsu M (2005) Bioorg Med Chem 13:4721–4732
Fujikawa M, Nakao K, Shimizu R, Akamatsu M (2007) Bioorg Med Chem 15:3756–3767
Verma RP, Hansch C, Selassie CD (2007) J Comput Aided Mol Des 21:3–22
Sun H, Nguyen K, Kerns E, Yan Z, Yu KR, Shah P, Jadhav A, Xu X (2017) Bioorg Med Chem 25:1266–1276
Oja M, Maran U (2015) Mol Inform 34:493–506
Oja M, Maran U (2016) SAR QSAR Environ Res 27:813–832
Oja M, Maran U (2018) Eur J Pharm Sci 123:429–440
Avdeef A, Tsinman O (2006) Eur J Pharm Sci 28:43–50
Diukendjieva A, Alov A, Tsakovska I, Pencheva T, Richarz A, Kren V, Cronin MTD, Pajeva I (2019) Phytomedicine 53:79–85
Karleson M, Karleson G, Tamm T, Tulp I, Jänes J, Tämm K, Lomaka A, Savchenko D, Dobchev A (2009) ARKIVOC Online J Org Chem 2009:218–238
Hansen N, van Gunsteren WF (2014) J Chem Theory Comput 10:2632–2647
Skyner RE, McDonagh JL, Groom CR, van Mourik T, Mitchell JBO (2015) Phys Chem Chem Phys 17:6174–6191
Chandler D, McCoy JD, Singer SJ (1986) J Chem Phys 85:5971–5976
Chandler D, McCoy JD, Singer SJ (1986) J Chem Phys 85:5977–5982
Lowden LJ, Chandler D (1973) J Chem Phys 59:6587–6595
Kovalenko A (2015) Condens Matter Phys 18:32601
Palmer DS, Frolov A, Ratkova EL, Fedorov MV (2010) J Phys Condens Matter 22:492101
Kovalenko A, Hirata F (2005) Phys Chem Chem Phys 7:1785–1793
Kovalenko A (2013) Pure Appl Chem 85:159–199
Kovalenko A (2017) Multiscale modeling of solvation. In: Breitkopf C, Swider-Lyons K (eds) Springer handbook of electrochemical energy. Springer, Berlin
Kovalenko A, Gusarov S (2017) Phys Chem Chem Phys 20:2947–2969
Ratkova EL, Palmer DS, Fedorov MV (2015) Chem Rev 13:6312–6356
Roy D, Kovalenko A (2019) J Phys Chem A 123:4087–4093
Roy D, Blinov N, Kovalenko A (2017) J Phys Chem B 121:9268–9273
Roy D, Hinge VK, Kovalenko A (2019) ACS Omega 4:3055–3060
Roy D, Hinge VK, Kovalenko A (2019) ACS Omega 4:16774–16780
Roy D, Kovalenko A (2019) J Comput Aided Mol Des 33:905–912
Roy D, Kovalenko A (2020) J Phys Chem B 124:4590–4597
Truchon J-F, Pettit BM, Labute P (2014) J Chem Theory Comput 10:934–941
Misin M, Palmer DS, Fedorov MV (2016) J Phys Chem B 2016(120):5724–5731
Avdeef A (2002) In: van de Waterbeemd H, Lennernas H, Artursson P (eds) Drug bioavailability. Weinheim, Wiley-VCH
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) J Cheminform 3:33
Zhao Y, Truhlar DG (2008) Theor Chem Acc 120:215–241
Easton RE, Geisen DJ, Welch A, Cramer CJ, Truhlar DG (1996) Theor Chem Acc 93:281–301
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Petersson GA, Nakatsuji H, Li X, Caricato M, Marenich AV, Bloino J, Janesko BG, Gomperts R, Mennucci B, Hratchian HP, Ortiz JV, Izmaylov AF, Sonnenberg JL, Williams-Young D, Ding F, Lipparini F, Egidi F, Goings J, Peng B, Petrone A, Henderson T, Ranasinghe D, Zakrzewski VG, Gao J, Rega N, Zheng G, Liang W, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Throssell K, Montgomery JA Jr., Peralta JE, Ogliaro F, Bearpark MJ, Heyd JJ, Brothers EN, Kudin KN, Staroverov VN, Keith TA, Kobayashi R, Normand J, Raghavachari K, Rendell AP, Burant JC, Iyengar SS, Tomasi J, Cossi M, Millam JM, Klene M, Adamo C, Cammi R, Ochterski JW, Martin RL, Morokuma K, Farkas O, Foresman JB, Fox DJ (2016) Gaussian 16, Revision B.01. Gaussian, Inc., Wallingford CT
Case DA, Ben-Shalom IY, Brozell SR, Cerutti DS, Cheatham TE III, Cruzeiro VWD, Darden TA, Duke RE, Ghoreishi D, Gilson MK, Gohlke H, Goetz AW, Greene D, Harris R, Homeyer N, Izadi S, Kovalenko A, Kurtzman T, Lee TS, LeGrand S, Li P, Lin C, Liu J, Luchko T, Luo R, Mermelstein DJ, Merz KM, Miao Y, Monard G, Nguyen C, Nguyen H, Omelyan I, Onufriev A, Pan F, Qi R, Roe DR, Roitberg A, Sagui C, Schott-Verdugo S, Shen J, Simmerling CL, Smith J, Salomon-Ferrer R, Swails J, Walker RC, Wang J, Wei H, Wolf RM, Wu X, Xiao L, York DM, Kollman PA (2018) AMBER 2018. University of California, San Francisco
Rappe AK, Casewit CJ, Colwell KS, Goddard WA, Skiff WM (1992) J Am Chem Soc 114:10024–10035
Dewar MJS, Zoebisch EG, Healy EF, Stewart JJP (1985) J Am Chem Soc 107:3902–3909
MOPAC2016, Stewart JJP, Stewart Computational Chemistry, Colorado Springs, CO, USA
Yap CW (2011) J Comput Chem 32:1466–1476
Marvin 17.21.0, ChemAxon (https://www.chemaxon.com)
R Core Team (2017) R: A Language And Environment For Statistical Computing; R Foundation for Statistical Computing, Vienna, Austria
Robinson D, Gomez M, Demeshev B, Menne D, Nutter B, Johnston L, Bolker B, Briatte F, Arnold J, Gabry J (2017) Broom: Convert Statistical Analysis Objects into Tidy Data Frames
Wickham H (2007) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York
Kuhn M (2008) Building predictive models in R using the caret package. J Stat Softw 1(5):2008
Ridgeway G (2007) Generalized boosted models: a guide to the gbm package. R package vignette, http://CRAN.R-project.org/package=gbm. Accessed 12 May 2020
Liaw R, Wiener M (2002) Classification and regression by randomForest. R News 2:18–22
Dimitriadou E, Hornik K, Leisch F, Meyer D, Weingessel A (2008) e1071: Misc Functions of the Department of Statistics (e1071), TU Wien. R package version 1.5-18. https://cran.r-project.org/web/packages/e1071/index.html. Accessed 12 May 2020
Schliep K, Hechenbichler K (2016) Weighted k-nearest neighbors for classification, regression and clustering. R package version 1.3. https://cran.r-project.org/web/packages/kknn/index.html. Accessed 12 May 2020
Eric S, Kalinic M, Ilic K, Zloh M (2014) SAR QSAR Environ Res 25:939–966
Kuhn M, Johnson K (2018) Applied predictive modeling. Springer, New York
James G, Witten D, Hastie T, Tibshirani R (2014) An introduction to statistical learning with applications in R. Springer, New York
Natekin A, Knoll A (2013) Front Neurorobot 7:21
Ertl P, Rhode B, Selzer P (2000) J Med Chem 43:3714–3717
Acknowledgements
This work was financially supported by the NSERC Discovery Grant (RES0029477), Alberta Innovates AARP VII Research Grant (RES0043948), and Alzheimer Society of Alberta and Northwest Territories AARP VII Research Grant (RES0043949). Generous computing time provided by WestGrid (www.westgrid.ca) and Compute Canada/Calcul Canada (www.computecanada.ca) is acknowledged. The authors thank Dr. Marcia LeVatte for assistance in editing the manuscript.
Author information
Authors and Affiliations
Contributions
All the authors contributed equally to writing the manuscript and approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Roy, D., Dutta, D., Wishart, D.S. et al. Predicting PAMPA permeability using the 3D-RISM-KH theory: are we there yet?. J Comput Aided Mol Des 35, 261–269 (2021). https://doi.org/10.1007/s10822-020-00364-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-020-00364-4