Abstract
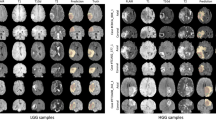
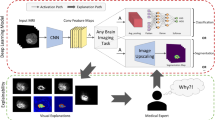
Precise neurosurgical guidance is critical for successful brain surgeries and plays a vital role in all phases of image-guided neurosurgery (IGN). Neuronavigation software enables real-time tracking of surgical tools, ensuring their presentation with high precision in relation to a virtual patient model. Therefore, this work focuses on the development of a novel multimodal IGN system, leveraging deep learning and explainable AI to enhance brain tumor surgery outcomes. The study establishes the clinical and technical requirements of the system for brain tumor surgeries. NeuroIGN adopts a modular architecture, including brain tumor segmentation, patient registration, and explainable output prediction, and integrates open-source packages into an interactive neuronavigational display. The NeuroIGN system components underwent validation and evaluation in both laboratory and simulated operating room (OR) settings. Experimental results demonstrated its accuracy in tumor segmentation and the success of ExplainAI in increasing the trust of medical professionals in deep learning. The proposed system was successfully assembled and set up within 11 min in a pre-clinical OR setting with a tracking accuracy of 0.5 (± 0.1) mm. NeuroIGN was also evaluated as highly useful, with a high frame rate (19 FPS) and real-time ultrasound imaging capabilities. In conclusion, this paper describes not only the development of an open-source multimodal IGN system but also demonstrates the innovative application of deep learning and explainable AI algorithms in enhancing neuronavigation for brain tumor surgeries. By seamlessly integrating pre- and intra-operative patient image data with cutting-edge interventional devices, our experiments underscore the potential for deep learning models to improve the surgical treatment of brain tumors and long-term post-operative outcomes.






Similar content being viewed by others
Data availability
The data used in this study were obtained from the BraTS Challenge 2021 dataset, which is publicly available. The details on accessing the BraTS dataset can be found on the official BraTS Challenge website (https://www.synapse.org/#!Synapse:syn25829067/). The software components developed in this study, including NeuroIGN, will be made available as open source and can be accessed through https://github.com/razeineldin/NeuroIGN.
Notes
Simplified user interface (Slicelet); https://www.slicer.org/wiki/Documentation/Nightly/Developers/Slicelets/.
NeuroIGN project website; https://www.github.com/razeineldin/NeuroGN/.
References
Schipmann-Miletić S, Stummer W (2020) Image-Guided Brain Surgery. In: Molecular Imaging in Oncology. Recent Results in Cancer Research. pp 813–841. https://doi.org/10.1007/978-3-030-42618-7_26
Miner RC (2017) Image-Guided Neurosurgery. J Med Imaging Radiat Sci 48 (4):328–335. https://doi.org/10.1016/j.jmir.2017.06.005
Sanai N, Polley MY, McDermott MW, Parsa AT, Berger MS (2011) An extent of resection threshold for newly diagnosed glioblastomas. J Neurosurg 115 (1):3–8. https://doi.org/10.3171/2011.2.jns10998
Hervey-Jumper SL, Berger MS (2016) Maximizing safe resection of low- and high-grade glioma. J Neurooncol 130 (2):269–282. https://doi.org/10.1007/s11060-016-2110-4
Fedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin JC et al. (2012) 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn Reson Imaging 30 (9):1323–1341. https://doi.org/10.1016/j.mri.2012.05.001
Goch CJ, Metzger J, Nolden M (2017) Abstract: Medical Research Data Management Using MITK and XNAT. In: Bildverarbeitung für die Medizin 2017. Informatik aktuell. pp 305–305. https://doi.org/10.1007/978-3-662-54345-0_68
Yushkevich PA, Piven J, Hazlett HC, Smith RG, Ho S et al. (2006) User-guided 3D active contour segmentation of anatomical structures: Significantly improved efficiency and reliability. NeuroImage 31 (3):1116–1128. https://doi.org/10.1016/j.neuroimage.2006.01.015
Ungi T, Lasso A, Fichtinger G (2016) Open-source platforms for navigated image-guided interventions. Med Image Anal 33:181–186. https://doi.org/10.1016/j.media.2016.06.011
Clarkson MJ, Zombori G, Thompson S, Totz J, Song Y et al. (2015) The NifTK software platform for image-guided interventions: platform overview and NiftyLink messaging. Int J Comput Assist Radiol Surg 10 (3):301–316. https://doi.org/10.1007/s11548-014-1124-7
Tokuda J, Fischer GS, Papademetris X, Yaniv Z, Ibanez L et al. (2009) OpenIGTLink: an open network protocol for image-guided therapy environment. Int J Med Robot 5 (4):423–434. https://doi.org/10.1002/rcs.274
Lasso A, Heffter T, Rankin A, Pinter C, Ungi T et al. (2014) PLUS: open-source toolkit for ultrasound-guided intervention systems. IEEE Trans Biomed Eng 61 (10):2527–2537. https://doi.org/10.1109/TBME.2014.2322864
Drouin S, Kochanowska A, Kersten-Oertel M, Gerard IJ, Zelmann R et al. (2017) IBIS: an OR ready open-source platform for image-guided neurosurgery. Int J Comput Assist Radiol Surg 12 (3):363–378. https://doi.org/10.1007/s11548-016-1478-0
Askeland C, Solberg OV, Bakeng JB, Reinertsen I, Tangen GA et al. (2016) CustusX: an open-source research platform for image-guided therapy. Int J Comput Assist Radiol Surg 11 (4):505–519. https://doi.org/10.1007/s11548-015-1292-0
Serte S, Serener A, Al-Turjman F (2020) Deep learning in medical imaging: A brief review. Transactions on Emerging Telecommunications Technologies 33 (10). https://doi.org/10.1002/ett.4080
Chen X, Wang X, Zhang K, Fung K-M, Thai TC et al. (2022) Recent advances and clinical applications of deep learning in medical image analysis. Medical Image Analysis 79. https://doi.org/10.1016/j.media.2022.102444
Kv R, Prasad K, Peralam Yegneswaran P (2023) Segmentation and Classification Approaches of Clinically Relevant Curvilinear Structures: A Review. Journal of Medical Systems 47 (1). https://doi.org/10.1007/s10916-023-01927-2.
Bierbrier J, Eskandari M, Di Giovanni DA, Collins DL (2023) Towards Estimating MRI-Ultrasound Registration Error in Image-Guided Neurosurgery. IEEE Transactions on Ultrasonics, Ferroelectrics, and Frequency Control:1–1. https://doi.org/10.1109/tuffc.2023.3239320
Pirhadi A, Salari S, Ahmad MO, Rivaz H, Xiao Y (2022) Robust landmark-based brain shift correction with a Siamese neural network in ultrasound-guided brain tumor resection. International Journal of Computer Assisted Radiology and Surgery. https://doi.org/10.1007/s11548-022-02770-5
Zeineldin RA, Karar ME, Elshaer Z, Coburger J, Wirtz CR et al. (2022) Explainability of deep neural networks for MRI analysis of brain tumors. Int J Comput Assist Radiol Surg 17 (9):1673–1683. https://doi.org/10.1007/s11548-022-02619-x
van der Velden BHM, Kuijf HJ, Gilhuijs KGA, Viergever MA (2022) Explainable artificial intelligence (XAI) in deep learning-based medical image analysis. Medical Image Analysis 79. https://doi.org/10.1016/j.media.2022.102470.
Lalithadevi B, Krishnaveni S, Gnanadurai JSC (2023) A Feasibility Study of Diabetic Retinopathy Detection in Type II Diabetic Patients Based on Explainable Artificial Intelligence. Journal of Medical Systems 47 (1). https://doi.org/10.1007/s10916-023-01976-7.
Pesapane F, Volonte C, Codari M, Sardanelli F (2018) Artificial intelligence as a medical device in radiology: ethical and regulatory issues in Europe and the United States. Insights Imaging 9 (5):745–753. https://doi.org/10.1007/s13244-018-0645-y
Zeineldin RA, Karar ME, Coburger J, Wirtz CR, Burgert O (2020) DeepSeg: deep neural network framework for automatic brain tumor segmentation using magnetic resonance FLAIR images. Int J Comput Assist Radiol Surg 15 (6):909–920. https://doi.org/10.1007/s11548-020-02186-z
Zeineldin RA, Karar ME, Burgert O, Mathis-Ullrich F (2022) Multimodal CNN Networks for Brain Tumor Segmentation in MRI: A BraTS 2022 Challenge Solution.arXiv:2212.09310. https://doi.org/10.48550/arXiv.2212.09310
Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K et al. (2015) The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS). IEEE Trans Med Imaging 34 (10):1993–2024. https://doi.org/10.1109/TMI.2014.2377694
Bakas S, Akbari H, Sotiras A, Bilello M, Rozycki M et al. (2017) Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci Data 4 (1):170117. https://doi.org/10.1038/sdata.2017.117
Bakas S, Reyes M, Jakab A, Bauer S, Rempfler M et al. (2018) Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv preprint arXiv:181102629
Baid U, Ghodasara S, Bilello M, Mohan S, Calabrese E et al. (2021) The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification. https://arxiv.org/abs/2107.02314
Shapey J, Dowrick T, Delaunay R, Mackle EC, Thompson S et al. (2021) Integrated multi-modality image-guided navigation for neurosurgery: open-source software platform using state-of-the-art clinical hardware. International Journal of Computer Assisted Radiology and Surgery 16 (8):1347–1356. https://doi.org/10.1007/s11548-021-02374-5
Acknowledgements
The authors would like to acknowledge the valuable contributions of the neurosurgeons and medical staff from the Department of Neurosurgery at the University Hospital Ulm/Günzburg for their expertise and support throughout the development and evaluation of the proposed AI-driven system for neurosurgery.
Funding
The first author was financially supported during this work by the German Academic Exchange Service (DAAD) [scholarship number 91705803].
Author information
Authors and Affiliations
Contributions
R.A.Z. and M.E.K. conceived the study and secured funding. R.A.Z. conducted investigations, developed the methodology, implemented the software, created visualizations, and drafted the original manuscript. M.E.K. provided supervision, contributed to methodology, investigations, and reviewed and edited the manuscript. O.B. contributed to the conceptualization, funding acquisition, investigations, resource management, and provided supervision for validation. F.MU. contributed to the conceptualization, investigations, resource management, and supervision for validation. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Ethical approval
As the data used in this study were obtained from a publicly available dataset, specifically the BraTS Challenge 2021, ethical approval was previously obtained by the BraTS organizers. The study followed the guidelines and protocols established by the BraTS Challenge for data usage.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zeineldin, R.A., Karar, M.E., Burgert, O. et al. NeuroIGN: Explainable Multimodal Image-Guided System for Precise Brain Tumor Surgery. J Med Syst 48, 25 (2024). https://doi.org/10.1007/s10916-024-02037-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10916-024-02037-3