Abstract
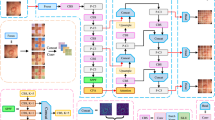
Small target polyps are prone to missed detection due to their small coverage area and little information. To address this issue, a modified PATM-YOLO polyp detection model based on YOLOv5 is proposed. The model first addresses the issue of missed detection of small polyps by constructing a detection head for identifying small polyps and using an improved Phase-Aware Token Mixing Module(PATM) attention module to increase the network’s attention to small polyps and suppress the model’s focus on non-polyp regions. Secondly, an improved Adaptively Spatial Feature Fusion(ASFF) module is proposed to fully utilize multi-scale information, enhancing the network’s feature richness. Finally, by introducing the Swin Transformer into the network and determining its optimal placement through experiments, the detection accuracy is maximized without affecting the network’s performance. After experimental comparison on the constructed dataset and the public dataset SUN, the proposed PATM-YOLO network model alleviated missed detection in dense and small polyp images, and achieved an precision of 91.3\(\%\), which is 8.5\(\%\) higher than the baseline YOLOv5 network model. This indicates that the detection performance of this model outperforms other classical target detection networks and the original network.









Similar content being viewed by others
Code Availibility
Not applicable
Availability of data and materials
Three publicly available polyp datasets of endoscopy images (the PLoS One-Zhang dataset, the Hyper-Kvasir-Segmentation dataset, and the SUN dataset) were used in the experiments of this study. The PLoS One-Zhang dataset can be found at: https://github.com/jiquan/Dataset-acess-for-PLOS-ONE. The Hyper-Kvasir-Segmentation dataset can be found at: https://datasets.simula.no/hyper-kvasir/. The SUN dataset can be found at: http://sundatabase.org.
References
Brosens LA, Wood LD, Offerhaus GJ, Arnold CA, Lam-Himlin D, Giardiello FM, Montgomery EA (2016) Pathology and genetics of syndromic gastric polyps. International journal of surgical pathology 24(3):185–199. https://doi.org/10.1177/1066896915620013
Ameling S, Wirth S, Paulus D, Lacey G, Vilarino F (2009) Texture-based polyp detection in colonoscopy. In: Bildverarbeitung Für die Medizin 2009: Algorithmen—Systeme—Anwendungen Proceedings des Workshops Vom 22. Bis 25. März 2009 in Heidelberg, Springer, pp 346–350. https://doi.org/10.1007/978-3-540-93860-6_70
Iakovidis DK, Maroulis DE, Karkanis SA, Brokos A (2005) A comparative study of texture features for the discrimination of gastric polyps in endoscopic video. In: 18th IEEE Symposium on computer-based medical systems (CBMS’05), IEEE, pp 575–580. https://doi.org/10.1109/CBMS.2005.6
Hwang S, Oh J, Tavanapong W, Wong J, De Groen PC (2007) Polyp detection in colonoscopy video using elliptical shape feature. In: 2007 IEEE International conference on image processing, IEEE, 2:465. https://doi.org/10.1109/ICIP.2007.4379193
Bernal J, Sánchez J, Vilarino F (2012) Towards automatic polyp detection with a polyp appearance model. Pattern Recognition 45(9):3166–3182. https://doi.org/10.1016/j.patcog.2012.03.002
Leung WK, Guo C-G, Ko MK, To EW, Mak LY, Tong TS, Chen L-J, But DY, Wong SY, Liu KS et al (2020) Linked color imaging versus narrow-band imaging for colorectal polyp detection: a prospective randomized tandem colonoscopy study. Gastrointestinal Endoscopy 91(1):104–112. https://doi.org/10.1016/j.gie.2019.06.031
Alexandre LA, Nobre N, Casteleiro J (2008) Color and position versus texture features for endoscopic polyp detection. In: 2008 International conference on biomedical engineering and informatics, IEEE, 2:38–42. https://doi.org/10.1109/BMEI.2008.246
Freedman JS, Harari DY, Bamji ND, Bodian CA, Kornacki S, Cohen LB, Miller KM, Aisenberg J (2011) The detection of premalignant colon polyps during colonoscopy is stable throughout the workday. Gastrointestinal endoscopy 73(6):1197–1206. https://doi.org/10.1016/j.gie.2011.01.019
Simmons DT, Harewood GC, Baron TH, Petersen BT, Wang KK, Boyd-Enders F, Ott BJ (2006) Impact of endoscopist withdrawal speed on polyp yield: implications for optimal colonoscopy withdrawal time. Alimentary pharmacology & therapeutics 24(6):965–971. https://doi.org/10.1016/j.gie.2006.03.026
Setio AAA, Ciompi F, Litjens G, Gerke P, Jacobs C, Van Riel SJ, Wille MMW, Naqibullah M, Sánchez CI, Van Ginneken B (2016) Pulmonary nodule detection in ct images: false positive reduction using multi-view convolutional networks. IEEE transactions on medical imaging 35(5):1160–1169. https://doi.org/10.1109/TMI.2016.2536809
Pang S, Ding T, Qiao S, Meng F, Wang S, Li P, Wang X (2019) A novel yolov3-arch model for identifying cholelithiasis and classifying gallstones on ct images. PloS one 14(6):0217647. https://doi.org/10.1371/journal.pone.0217647
Deeba F, Bui FM, Wahid KA (2020) Computer-aided polyp detection based on image enhancement and saliency-based selection. Biomed Signal Process Control 55:101530. https://doi.org/10.1016/j.bspc.2019.04.007
Qadir HA, Shin Y, Solhusvik J, Bergsland J, Aabakken L, Balasingham I (2021) Toward real-time polyp detection using fully cnns for 2d gaussian shapes prediction. Med Image Anal 68:101897. https://doi.org/10.1016/j.media.2020.101897
Taş M, Yılmaz B (2021) Super resolution convolutional neural network based pre-processing for automatic polyp detection in colonoscopy images. Comput Electrical Eng 90:106959. https://doi.org/10.1016/j.compeleceng.2020.106959
Chen B-L, Wan J-J, Chen T-Y, Yu Y-T, Ji M (2021) A self-attention based faster r-cnn for polyp detection from colonoscopy images. Biomed Signal Process Control 70:103019. https://doi.org/10.1016/j.bspc.2021.103019
Cao C, Wang R, Yu Y, Zhang H, Yu Y, Sun C (2021) Gastric polyp detection in gastroscopic images using deep neural network. PloS one 16(4):0250632. https://doi.org/10.1371/journal.pone.0250632
Nisha J, Gopi VP, Palanisamy P (2022) Automated colorectal polyp detection based on image enhancement and dual-path cnn architecture. Biomed Signal Process Control 73:103465. https://doi.org/10.1016/j.bspc.2021.103465
Hu K, Zhao L, Feng S, Zhang S, Zhou Q, Gao X, Guo Y (2022) Colorectal polyp region extraction using saliency detection network with neutrosophic enhancement. Comput Biol Med 147:105760. https://doi.org/10.1016/j.compbiomed.2022.105760
Zhu X, Lyu S, Wang X, Zhao Q (2021) Tph-yolov5: Improved yolov5 based on transformer prediction head for object detection on drone-captured scenarios. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 2778–2788. https://doi.org/10.48550/arXiv.2108.11539
Tang Y, Han K, Guo J, Xu C, Li Y, Xu C, Wang Y (2022) An image patch is a wave: Phase-aware vision mlp. In: Proceedings of the IEEE/CVF Conference on computer vision and pattern recognition, pp 10935–10944. https://doi.org/10.48550/arXiv.2111.12294
Liu Z, Hu H, Lin Y, Yao Z, Xie Z, Wei Y, Ning J, Cao Y, Zhang Z, Dong L, et al. (2022) Swin transformer v2: Scaling up capacity and resolution. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 12009–12019. https://doi.org/10.48550/arXiv.2111.09883
Liu S, Huang D, Wang Y (2019) Learning spatial fusion for single-shot object detection. arXiv preprint arXiv:1911.09516. https://doi.org/10.48550/arXiv.1911.09516
Zhang X, Chen F, Yu T, An J, Huang Z, Liu J, Hu W, Wang L, Duan H, Si J (2019) Real-time gastric polyp detection using convolutional neural networks. PloS one 14(3):0214133. https://doi.org/10.1371/journal.pone.0214133
Borgli H, Thambawita V, Smedsrud PH, Hicks S, Jha D, Eskeland SL, Randel KR, Pogorelov K, Lux M, Nguyen DTD et al (2020) Hyperkvasir, a comprehensive multi-class image and video dataset for gastrointestinal endoscopy. Scientific data 7(1):283. https://doi.org/10.1038/s41597-020-00622-y
Misawa M, Kudo S-E, Mori Y, Hotta K, Ohtsuka K, Matsuda T, Saito S, Kudo T, Baba T, Ishida F et al (2021) Development of a computer-aided detection system for colonoscopy and a publicly accessible large colonoscopy video database (with video). Gastrointestinal endoscopy 93(4):960–967. https://doi.org/10.1016/j.gie.2020.07.060
Pacal I, Karaman A, Karaboga D, Akay B, Basturk A, Nalbantoglu U, Coskun S (2022) An efficient real-time colonic polyp detection with yolo algorithms trained by using negative samples and large datasets. Comput Biol Med 141:105031. https://doi.org/10.1016/j.compbiomed.2021.105031
Karaman A, Pacal I, Basturk A, Akay B, Nalbantoglu U, Coskun S, Sahin O, Karaboga D (2023) Robust real-time polyp detection system design based on yolo algorithms by optimizing activation functions and hyper-parameters with artificial bee colony (abc). Expert Syst Appl 221:119741. https://doi.org/10.1016/j.eswa.2023.119741
Funding
This work was supported by the Shanghai Science and Technology Innovation Action Plan(22S31903700 \( \& \) 21S31904200).
Author information
Authors and Affiliations
Contributions
LW, JL, and HY contributed to conception and design of the study. LW, HY, HL, and SC organized the database. LW, JL and SC performed the statistical analysis. LW wrote the manuscript. All authors contributed to manuscript revision, read, and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Ethics approval
Not applicable
Consent to participate
Not applicable
Consent for publication
Not applicable
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, L., Liu, J., Yang, H. et al. Small gastric polyp detection based on the improved YOLOv5. Multimed Tools Appl 83, 71773–71788 (2024). https://doi.org/10.1007/s11042-024-18497-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-024-18497-1