Abstract
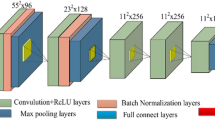
The accurate classification of the histopathological images of breast cancer diagnosis may face a huge challenge due to the complexity of the pathologist images. Currently, computer-aided diagnosis is implemented to get sound and error-less diagnosis of this lethal disease. However, the classification accuracy and processing time can be further improved. This study was designed to control diagnosis error via enhancing image accuracy and reducing processing time by applying several algorithms such as deep learning, K-means, autoencoder in clustering and enhanced loss function (ELF) in classification. Histopathological images were obtained from five datasets and pre-processed by using stain normalisation and linear transformation filter. These images were patched in sizes of 512 × 512 and 128 × 128 and extracted to preserve the tissue and cell levels to have important information of these images. The patches were further pre-trained by ResNet50-128 and ResNet512. Meanwhile, the 128 × 128 were clustered and autoencoder was employed with K-means which used latent feature of image to obtain better clustering result. Classification algorithm is used in current proposed system to ELF. This was achieved by combining SVM loss function and optimisation problem. The current study has shown that the deep learning algorithm has increased the accuracy of breast cancer classification up to 97% compared to state-of-the-art model which gave a percentage of 95%, and the time was decreased to vary from 30 to 40 s. Also, this work has enhanced system performance via improving clustering by employing K-means with autoencoder for the nonlinear transformation of histopathological image.




Similar content being viewed by others
References
Li Y, Wu J, Wu Q (2019) Classification of breast cancer histology images using multi-size and discriminative patches based on deep learning. IEEE Access 7:21400–21408
Vo DM, Nguyen N-Q, Lee S-W (2019) Classification of breast cancer histology images using incremental boosting convolution networks. Inf Sci 482:123–138
Jadoon MM, Zhang Q, Haq IU, Butt S, Jadoon A (2017) Three-class mammogram classification based on descriptive cnn features. BioMed Res Int 3640901:11. https://doi.org/10.1155/2017/3640901
Wahab N, Khan A, Lee YS (2017) Two-phase deep convolutional neural network for reducing class skewness in histopathological images based breast cancer detection. Comput Bio Med 85:86–97
Xie J, Liu R, Luttrell J, Zhang C (2019) Deep learning based analysis of histopathological images of breast cancer. Front Genet 10:80
Mambou SJ, Maresova P, Krejcar O, Selamat A, Kuca K (2018) breast cancer detection using infrared thermal imaging and a deep learning model, (in eng). Sensors (Basel, Switzerland) 18(9):2799
Chougrad H, Zouaki H, Alheyane O (2018) Deep convolutional neural networks for breast cancer screening. Comput Methods Programs Biomed 157:19–30
Nahid A-A, Mehrabi AM, Kong Y (2018) Histopathological breast cancer image classification by deep neural network techniques guided by local clustering. Biomed Res Int 03:07
Spanhol FA, Oliveira LS, Petitjean C, Heutte L (2016) Breast cancer histopathological image classification using convolutional neural networks. In: 2016 International joint Conference on neural networks (IJCNN), pp 2560–2567
Roy K, Banik D, Bhattacharjee D, Nasipuri M (2019) Patch-based system for classification of breast histology images using deep learning. Comput Med Imaging Graph 71:90–103
Gandomkar Z, Brennan PC, Mello-Thoms C (2018) MuDeRN: multi-category classification of breast histopathological image using deep residual networks. Artif Intell Med 88:14–24
Zheng Y et al (2017) Feature extraction from histopathological images based on nucleus-guided convolutional neural network for breast lesion classification. Pattern Recogn 71:14–25
Jiao Z, Gao X, Wang Y, Li J (2016) A deep feature based framework for breast masses classification. Neurocomputing 197:221–231
Han Z, Wei B, Zheng Y, Yin Y, Li K, Li S (2017) Breast cancer multi-classification from histopathological images with structured deep learning model. Sci Rep 7(1):4172
Sudharshan PJ, Petitjean C, Spanhol F, Oliveira LE, Heutte L, Honeine P (2019) Multiple instance learning for histopathological breast cancer image classification. Exp Syst Appl 117:103–111
Hamidinekoo A, Denton E, Rampun A, Honnor K, Zwiggelaar R (2018) Deep learning in mammography and breast histology, an overview and future trends. Med Image Anal 47:45–67
Araújo T et al (2017) Classification of breast cancer histology images using convolutional neural networks. PLoS ONE 12(6):e0177544
Suzuki S, et al (2016) Mass detection using deep convolutional neural network for mammographic computer-aided diagnosis. In: 2016 55th Annual Conference of the Society of Instrument and Control Engineers of Japan (SICE), pp 1382–1386
Ragab DA, Sharkas M, Marshall S, Ren J (2019) Breast cancer detection using deep convolutional neural networks and support vector machines. PeerJ 7:e6201
Mohamed AA, Berg WA, Peng H, Luo Y, Jankowitz RC, Wu S (2018) A deep learning method for classifying mammographic breast density categories. Med Phys 45(1):314–321
Al-masni MA et al (2018) Simultaneous detection and classification of breast masses in digital mammograms via a deep learning YOLO-based CAD system. Comput Methods Programs Biomed 157:85–94
Asri H, Mousannif H, Moatassime HA, Noel T (2016) Using machine learning algorithms for breast cancer risk prediction and diagnosis. Procedia Comput Sci 83:1064–1069
Motlagh MH et al (2018) Breast cancer histopathological image classification: a deep learning approach. bioRxiv, p 1–8. https://doi.org/10.1109/BIBM.2018.8621307
Ting FF, Tan YJ, Sim KS (2019) Convolutional neural network improvement for breast cancer classification. Expert Syst Appl 120:103–115
Quellec G, Cazuguel G, Cochener B, Lamard M (2017) Multiple-instance learning for medical image and video analysis. IEEE Rev Biomed Eng 10:213–234
Funding
There was no grant support for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors indicate full freedom of investigation and no potential conflict of interest.
Ethical approval
Not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Acharya, S., Alsadoon, A., Prasad, P.W.C. et al. Deep convolutional network for breast cancer classification: enhanced loss function (ELF). J Supercomput 76, 8548–8565 (2020). https://doi.org/10.1007/s11227-020-03157-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11227-020-03157-6