Abstract
In the present paper, deep convolutional neural network (DCNN) is applied to multilocus protein subcellular localization as it is more suitable for multi-class classification. There are two main problems with this application. First, the appropriate features for correlation between multiple sites are hard to find. Second, the classifier structure is difficult to determine as it is greatly affected by the distribution of classified data. To solve these problems, a self-evoluting framework using DCNNs for multilocus protein subcellular localization is proposed. It has three characteristics that the previous algorithms do not. The first is that it combines the ant colony algorithm with the DCNN to form a self-evoluting algorithm for multilocus protein subcellular localization. The second is that it randomly groups subcellular sites using a limited random k-labelsets multi-label classification method. It also solves complex problems in a divide-and-conquer approach and proposes a flexible expansion model. The third is that it realizes the random selection feature extraction method in the positioning process and avoids the defects in individual feature extraction methods. The algorithm in the present paper is tested on the human database, and the overall correct rate is 67.17%, which is higher than that for the stacked self-encoder (SAE), support vector machine (SVM), random forest classifier (RF), or single deep convolutional neural network.
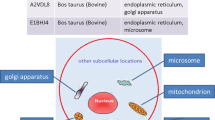
Graphical abstract

The algorithm mentioned in the present paper mainly includes four parts. They are protein sequence data preprocessing, integrated DCNN model construction, finding optimal DCNN combination by ant colony optimization, and protein subcellular localization for sequences. These parts are sequential relationships and the data obtained in the previous part is the basis for the latter part of the function. In the part of data preprocessing, the limited RAkEL multi-label classification method is used to randomly group subcellular sites. At the same time, the feature fusion of protein sequences is carried out by using multiple feature extraction methods. Each combination including features and sites information corresponds to a DCNN model. In the part of finding optimal DCNN combination by ant colony optimization, the main purpose is to find the best combination of DCNN models through the global optimization ability of the ant colony algorithm. The positioning of sequences is mainly to obtain multilocus subcellular localization by the optimal model combination.














Similar content being viewed by others
References
Yang L, Lv Y, Li T, Zuo Y, Jiang W (2014) Human proteins characterization with subcellular localizations. J Theor Biol 358:61–73
Ludwik KA, von Kuegelgen N, Chekulaeva M (2019) Genome-wide analysis of RNA and protein localization and local translation in mesc-derived neurons. Methods 162-163:31–41
Wei L, Liao M, Gao X, Wang J, Lin W (2016) mgof-loc: a novel ensemble learning method for human protein subcellular localization prediction. Neurocomputing 217:73–82
Mooney C, Wang Y-H, Pollastri G (2011) Sclpred: protein subcellular localization prediction by n-to-1 neural networks. Bioinformatics 27(20):2812–2819
Zhou H, Yang Y, Shen H-B (2016) Hum-mPLoc 3.0: prediction enhancement of human protein subcellular localization through modeling the hidden correlations of gene ontology and functional domain features. Bioinformatics 33(6):843–853
Wan S, Mak M-W, Kung S-Y (2016) Sparse regressions for predicting and interpreting subcellular localization of multi-label proteins. BMC Bioinf 17(1):97
Mak M-W, Guo J, Kung S-Y (2008) Pairprosvm: protein subcellular localization based on local pairwise profile alignment and svm. IEEE/ACM Trans Comput Biol Bioinform 5(3):416–422
Zhang S, Duan X (2018) Prediction of protein subcellular localization with oversampling approach and Chou’s general PseAAC. J Theor Biol 437:239–250
Zhao D, Liu H, Zheng Y, He Y, Lu D, Lyu C (2019) A reliable method for colorectal cancer prediction based on feature selection and support vector machine. Med Biol Eng Comput 57(4):901–912
Cheng X, Xiao X, Chou K-C (2017) pLoc-mHum: predict subcellular localization of multi-location human proteins via general PseAAC to winnow out the crucial GO information. Bioinformatics 34(9):1448–1456
Zhang D, Huang H, Bai X, Fang X, Zhang Y (2019) A highprecision hybrid algorithm for predicting eukaryotic protein subcellular localization. BioRxiv, page 620179
Liu Z, Jianjun H (2016) Mislocalization-related disease gene discovery using gene expression based computational protein localization prediction. Methods 93:119–127
Chou K-C (2013) Some remarks on predicting multi-label attributes in molecular biosystems. Mol BioSyst 9(6):1092–1100
Barracchia EP, Pio G, D'Elia D, Ceci M (2020) Prediction of new associations between ncRNAs and diseases exploiting multi-type hierarchical clustering. BMC Bioinf 21(1)
Jiang X, Zhao J, Qian W, Song W, Lin GN (2020) A generative adversarial network model for disease gene prediction with RNA-seq data. IEEE Access 8:37352–37360
Pio G, Ceci M, Prisciandaro F, Malerba D (2019) Exploiting causality in gene network reconstruction based on graph embedding. Mach Learn 109:1231–1279
Li Z, Zhu J, Xu X, Yao Y (2020) Rdense: a protein–RNA binding prediction model based on bidirectional recurrent neural network and densely connected convolutional networks. IEEE Access 8:14588–14605
Mignone P, Pio G, D'Elia D, Ceci M (2019) Exploiting transfer learning for the reconstruction of the human gene regulatory network. Bioinformatics 36(5):1553–1561
Li J, Li Z, Luo J, Yao Y (2020) ACNNT3: attention-CNN framework for prediction of sequence-based bacterial type III secreted effectors. Comput Math Methods Med 2020:1–7
Yang F, Xu Y-Y, Wang S-T, Shen H-B (2014) Image-based classification of protein subcellular location patterns in human reproductive tissue by ensemble learning global and local features. Neurocomputing 131:113–123
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521(7553):436
Wang K, Li K, Zhou L, Hu Y, Cheng Z, Liu J, Chen C (2019) Multiple convolutional neural networks for multivariate time series prediction. Neurocomputing 360:107–119
Alik N, Kurban OC, Yilmaz AR et al (2019) Large-scale offline signature recognition via deep neural networks and feature embedding [J]. Neurocomputing 359
Kleinkauf R, Houwaart T, Backofen R, Mann M (2015) AntaRNA-multiobjective inverse folding of pseudoknot RNA using ant-colony optimization. BMC Bioinf 16(1):389
Wu Y, Gong M, Ma W, Wang S (2019) High-order graph matching based on ant colony optimization. Neurocomputing 328:97–104
Nápoles G, Falcon R, Dikopoulou Z, Papageorgiou E, Bello R, Vanhoof K (2017) Weighted aggregation of partial rankings using ant colony optimization. Neurocomputing 250:109–120
Cheng X, Xiao X, Wu Z-c, Wang P, Lin W-z (2013) Swfoldrate: predicting protein folding rates from amino acid sequence with sliding window method. Proteins: Struct, Funct, Bioinf 81(1):140–148
Shen H-B, Chou K-C (2010) Virus-mPLoc: a fusion classifier for viral protein subcellular location prediction by incorporating multiple sites. J Biomol Struct Dyn 28(2):175–186
Mei S (2012) Multi-kernel transfer learning based on chou’s PseAAC formulation for protein submitochondria localization. J Theor Biol 293:121–130
Schmidhuber J (2015) Deep learning in neural networks: an overview [J]. Neural Netw 61:85–117
Rawat W, Wang Z (2017) Deep convolutional neural networks for image classification: a comprehensive review. Neural Comput 29(9):2352–2449
Sui X, Zheng Y, Wei B, Bi H, Wu J, Pan X, Yin Y, Zhang S (2017) Choroid segmentation from optical coherence tomography with graph-edge weights learned from deep convolutional neural networks. Neurocomputing 237:332–341
Lee C-Y, Gallagher PW, Tu Z (2016) Generalizing pooling functions in convolutional neural networks: mixed, gated, and tree. In: Artificial Intelligence and Statistics, pp 464–472
Wen M, Zhang Z, Niu S, Sha H, Yang R, Yun Y, Lu H (2017) Deep-learning-based drug–target interaction prediction. J Proteome Res 16(4):1401–1409
Reed M, Yiannakou A, Evering R (2014) An ant colony algorithm for the multi-compartment vehicle routing problem. Appl Soft Comput 15:169–176
Gao Y, Guan H, Qi Z, Yang H, Liang L (2013) A multi-objective ant colony system algorithm for virtual machine placement in cloud computing. J Comput Syst Sci 79(8):1230–1242
Rogai F, Manfredi C, Bocchi L (2016) Metaheuristics for specialization of a segmentation algorithm for ultrasound images. IEEE Trans Evol Comput 20(5):730–741
Boutet E, Lieberherr D, Tognolli M, Schneider M, Bairoch A (2007) Uniprotkb/swiss-prot. In: Plant bioinformatics. Springer, Berlin, pp 89–112
Jaramillo-Garzon J, Castellanos-Dominguez A et al (2015) Feature extraction by statistical contact potentials and wavelet transform for predicting subcellular localizations in gram negative bacterial proteins [J]. J Theor Biol 364:121–130
Lian J, Shi Y, Zhang Y, Jia W, Fan X, Zheng Y (2020) Revealing false positive features in epileptic EEG identification. Int J Neural Syst:2050017–2050017
Mandal M, Mukhopadhyay A, Maulik U (2015) Prediction of protein subcellular localization by incorporating multiobjective PSO-based feature subset selection into the general form of Chou’s PseAAC. Med Biol Eng Comput 53(4):331–344
Dorigo M, Birattari M (2010) Ant colony optimization. Springer, Berlin
Dorigo M, Stützle T (2006) The ant colony optimization metaheuristic: algorithms, applications, and advances. In: Handbook of Metaheuristics
LeyiWei YD, Su R, Tang J, Zou Q (2017) Prediction of human protein subcellular localization using deep learning. J Parallel Distrib Comput 117:212–217
Pärnamaa T, Parts L (2017) Accurate classification of protein subcellular localization from high-throughput microscopy images using deep learning. G3: Genes, Genomes, Genetics 7(5):1385–1392
Acknowledgments
This research was supported by the National Natural Science Foundation of China (61876102, 61472232) and the National Key Research and Development Program of China (No. 2016YFC0106000).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cong, H., Liu, H., Chen, Y. et al. Self-evoluting framework of deep convolutional neural network for multilocus protein subcellular localization. Med Biol Eng Comput 58, 3017–3038 (2020). https://doi.org/10.1007/s11517-020-02275-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-020-02275-w