Abstract
Imaging genetics research can explore the potential correlation between imaging and genomics. Most association analysis methods cannot effectively use the prior knowledge of the original data. In this respect, we add the prior knowledge of each original data to mine more effective biomarkers. The study of imaging genetics based on the sparse canonical correlation analysis (SCCA) is helpful to mine the potential biomarkers of neurological diseases. To improve the performance and interpretability of SCCA, we proposed a penalty method based on the autocorrelation matrix for discovering the possible biological mechanism between single nucleotide polymorphisms (SNP) variations and brain regions changes of Alzheimer’s disease (AD). The addition of the penalty allows the proposed algorithm to analyze the correlation between different modal features. The proposed algorithm obtains more biologically interpretable ROIs and SNPs that are significantly related to AD, which has better anti-noise performance. Compared with other SCCA-based algorithms (JCB-SCCA, JSNMNMF), the proposed algorithm can still maintain a stronger correlation with ground truth even when the noise is larger. Then, we put the regions of interest (ROI) selected by the three algorithms into the SVM classifier. The proposed algorithm has higher classification accuracy. Also, we use ridge regression with SNPs selected by three algorithms and four AD risk ROIs. The proposed algorithm has a smaller root mean square error (RMSE). It shows that proposed algorithm has a good ability in association recognition and feature selection. Furthermore, it selects important features more stably, improving the clinical diagnosis of new potential biomarkers.
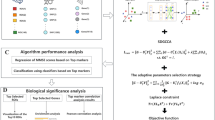
Graphical abstract








Similar content being viewed by others
Explore related subjects
Discover the latest articles and news from researchers in related subjects, suggested using machine learning.References
Thompson PM, Martin NG, Wright MJ (2010) Imaging genomics. Curr Opin Neurol 23(4):368–373
Hariri AR, Weinberger DR (2003) Imaging genomics[J]. Br Med Bull 65:259
Daniela M, Witten, et al (2009) Extensions of sparse canonical correlation analysis with applications to genomic data. Stat Appl Genet Molec Biol 8(1):1–27
Shen L, Risacher SL, Du L et al (2014) A novel structure-aware sparse learning algorithm for brain imaging genetics. Med Image Comput Comput Assist Interv 17(3):329–336
Christopher, Mark, Tang, Li, Fingert, John H, et al. Automated discovery of structural features of the optic nerve head on the basis of image and genetic data[J]. 2014.
Lin D, Calhoun VD, Wang YP (2014) Correspondence between fMRI and SNP data by group sparse canonical correlation analysis[J]. Med Image Anal 18(6):891–902
Hua W, Feiping N, Heng H et al (2012) From phenotype to genotype: an association study of longitudinal phenotypic markers to Alzheimer’s disease relevant SNPs[J]. Bioinformatics 28(18):619–625
Hua W, Feiping N, Heng H et al (2012) Identifying quantitative trait loci via group-sparse multitask regression and feature selection: an imaging genetics study of the ADNI cohort[J]. Bioinformatics 28(2):229–237
Vounou M, Koritakova E, Wolz R, Stein J, Thompson P, Rueckert D, Montana G (2011) Sparse reduced-rank regression detects genetic associations with voxel wise longitudinal phenotypes in Alzheimer’s disease. Neuroimage 60:700–716. https://doi.org/10.1016/j.neuroimage.2011.12.029
Meda SA, Narayanan B, Liu J et al (2012) A large scale multivariate parallel ICA method reveals novel imaging–genetic relationships for Alzheimer’s disease in the ADNI cohort[J]. Neuroimage 60(3):1608–1621
Du L, Yan J, Kim S, Risacher S, Huang H, Inlow M, Moore J, Saykin A, Shen L (2014) A novel structure-aware sparse learning algorithm for brain imaging genetics. Med Image Comput Assist Interv 17:329–336. https://doi.org/10.1007/978-3-319-10443-0_42
Du L, Liu K, Yao X, Risacher S, Han J, Saykin A, Guo L, Shen Li (2020) Detecting genetic associations with brain imaging phenotypes in Alzheimer’s disease via a novel structured SCCA approach. Med Image Anal 61:101656. https://doi.org/10.1016/j.media.2020.101656
Jingwen Y, Lei D, Sungeun K et al (2014) Transcriptome-guided amyloid imaging genetic analysis via a novel structured sparse learning algorithm[J]. Bioinformatics 30(17):564–571
Hao X, et al. (2017)“Mining Outcome-relevant Brain Imaging Genetic Associations via Three-way Sparse Canonical Correlation Analysis in Alzheimer's Disease.” Scientific reports, 7 p. 44272. https://doi.org/10.1038/srep44272
Kim M, Won JH, Youn J et al (2019) Joint-connectivity-based sparse canonical correlation analysis of imaging genetics for detecting biomarkers of Parkinson’s disease[J]. IEEE Trans Med Imaging 39(1):23–34
Fang J, Lin D, Schulz C et al (2016) Joint sparse canonical correlation analysis for detecting differential imaging genetics modules[J]. Bioinformatics 32:3480–3488
Daniela M (2009) Witten, Robert Tibshirani, Trevor Hastie, A penalized matrix decomposition, with applications to sparse principal components and canonical correlation analysis. Biostatistics 10(3):515–534
Tibshirani WRJ (2009) Extensions of sparse canonical correlation analysis with applications to genomic data[J]. Stat Appl Genet Molec Biol 8(1):28
Hoefling H (2010) A path algorithm for the fused lasso signal approximator[J]. J Comput Graph Stats 19(4):984–1006
Deng J, Zeng W, Kong W et al (2019) Multi-constrained joint non- negative matrix factorization with application to imaging genomic study of lung metastasis in soft tissue sarcomas[J]. IEEE Trans Biomed Eng 67(7):2110–2118
Knutson B (2013) Interpretable whole-brain prediction analysis with GraphNet[J]. Neuroimage 72(2):304–321
Lei Du, Heng, et al (2016) Structured sparse canonical correlation analysis for brain imaging genetics: an improved GraphNet method[J]. Bioinformatics 32(10):1544–1551
Angulo S, García-Pérez I, Legido-Quigley C, Barbas C (2009) The autocorrelation matrix probing biochemical relationships after metabolic fingerprinting with CE. Electrophoresis 30(7):1221–1227. https://doi.org/10.1002/elps.200800554
Gorski J, Pfeuffer F, Klamroth K (2007) Biconvex sets and optimization with biconvex functions: a survey and extensions[J]. Math Methods Oper Res 66(3):373–407
Xie S, Chen L, Zuo N et al (2016) DiffusionKit: A light one-stop solution for diffusion MRI data analysis[J]. J Neurosci Methods 273(273):107–119
Tate, David. (2017). Voxel-Based Morphometry. https://doi.org/10.1007/978-3-319-56782-2_9076-2.
Saykin AJ, Shen L, Foroud TM et al (2010) Alzheimer"s disease neuroimaging initiative biomarkers as quantitative phenotypes: genetics core aims, progress, and plans[J]. Alzheimers & Dementia the Journal of the Alzheimers Association 6(3):265–273
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: A tool set for whole-genome association and population-based linkage analyses[J]. Am J Hum Genet 81(3):559–575
Bertram L, Mcqueen MB, Mullin K et al (2007) Systematic meta-analyses of Alzheimer disease genetic association studies: the AlzGene database[J]. Nat Genet 39(1):17–23
Delaneau O, Zagury JF, Marchini J (2012) Improved whole-chromosome phasing for disease and population genetic studies[J]. Nat Methods 10(1):5–6
Jack CRJ, Petersen RC, Xu YC et al (1999) Prediction of AD with MRI- Based Hippocampal Volume in Mild Cognitive Impairment[J]. Neurology 52(7):1397–1403
Belmont DJW, Gibbs RA (2004) Genome-Wide Linkage Disequilibrium and Haplotype Maps[J]. Am J Pharmacogenomics 4(4):253–262
Barrett JC, Fry B, Maller J et al (2005) Haploview: analysis and visualization of LD and haplotype maps[J]. Bioinformatics 21(2):263–265
Kong V, Gabriel AD et al (2018) Early-in-life neuroanatomical and behavioural trajectories in a triple transgenic model of Alzheimer’s disease[J]. Brain Struct Funct 223(7):3365–3382
Jaroudi W, Garami J et al (2017) Factors underlying cognitive decline in old age and Alzheimer’s disease: the role of the hippocampus[J]. Rev Neurosci 28(7):705–714
Ertekin T, Acer N, Kseolu E et al (2016) Total intracranial and lateral ventricle volumes measurement in Alzheimer’s disease: a methodological study[J]. J Clin Neurosci 34:133–139
Grubman A, Chew G, Ouyang JF et al (2019) A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci 22:2087–2097. https://doi.org/10.1038/s41593-019-0539-4
Finger E, Zhang J, Dickerson B et al (2017) Disinhibition in Alzheimer’s disease is associated with reduced right frontal pole cortical thickness. J Alzheimer Dis 60(3):1161–1170
Snowden SG, Ebshiana AA, Hye A et al (2019) Neurotransmitter imbalance in the brain and Alzheimer’s disease pathology[J]. J Alzheimer Dis 72:1–9
Lin F, Ren P, Lo RY et al (2016) Insula and inferior frontal gyrus’ activities protect memory performance against Alzheimer’s disease pathology in old age[J]. J Alzheimer Dis 55(2):669–678
Kucukkilic E, Brookes K, Barber I et al (2018) Complement receptor 1 gene (CR1) intragenic duplication and risk of Alzheimer’s disease[J]. Hum Genet 137(1):1–10
Yonghong L, Andrew G, Charles R et al (2008) Evidence that common variation in NEDD9 is associated with susceptibility to late-onset Alzheimer’s and Parkinson’s disease[J]. Hum Mol Genet 17(5):759–767
Beck TN, Nicolas E, Kopp MC et al (2014) Adaptors for disorders of the brain? The cancer signaling proteins NEDD9, CASS4, and PTK2B in Alzheimer’s disease[J]. Oncoscience 1(7):486–503
(2019) Amyloidosis causes downregulation of SorLA, SorCS1 and SorCS3 expression in mice[J]. Biol Chem 400(9):1181-1189
Funding
This work was supported by the Natural Science Foundation of Shanghai (No. 18ZR1417200) and National Natural Science Foundation of China (No. 61803257).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wei, K., Kong, W. & Wang, S. Associating brain imaging phenotypes and genetic in Alzheimer’s disease via JSCCA approach with autocorrelation constraints. Med Biol Eng Comput 60, 95–108 (2022). https://doi.org/10.1007/s11517-021-02439-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-021-02439-2