Abstract
Dementia is a major cause of disability and dependency among older adults. Diagnosis is most effective at an early stage of the disease, as patients can start early treatment to delay progressive cognitive decline. While other diagnostic methods for dementia are available, electroencephalography (EEG) is noninvasive, more accessible, and less complicated than other biomarker measurements. Moreover, it may be orders of magnitude less expensive, thereby offering the possibility of low-cost mass screening. This paper presents a novel digital signal processing method called cardinal spline empirical mode decomposition (CS-EMD) for EEG processing. This new method uses a different signal envelope interpolation algorithm to separate EEG signals into constituent components, called intrinsic mode functions (IMFs), with better signal decomposition properties than classical empirical mode decomposition (EMD). The IMFs obtained from the new method are then used to compute longitudinal and transversal synchrony measures, which are explored as features for healthy and dementia classification using a support vector machine (SVM). The performance of the proposed method is studied on a publicly available EEG dataset. The results show that using multiple synchrony measures of both longitudinal and transversal EEG channels on five IMFs produces the best classification result of 90% accuracy, 96.67% specificity, 83.33% sensitivity, and 96.15% precision, outperforming the classical EMD method and various other approaches. This new, data-driven CS-EMD method shows good potential as a dementia screening tool. CS-EMD may also be applied in processing other nonlinear and nonstationary biosignals.
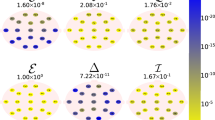
Graphical abstract






Similar content being viewed by others
References
Alzheimer’s Disease International (2015) The global impact of dementia: an analysis of prevalence, incidence, cost and trends. In: World Alzheimer report. https://www.alzint.org/resource/world-alzheimer-report-2015/
Waldemar G, Phungh KTT, Burns A, Georges J, Hansen FR, Iliffe S, Marking C, Olde-Rikkert M, Selmes J, Stoppe G, Sartorius N (2007) Access to diagnostic evaluation and treatment for dementia in Europe. Int J Geriatr Psychiatry 22(1):47–54. https://doi.org/10.1002/gps.1652
Prince MJ, Fan Wu, Guo Y, Gutierrez LM, Robledo MO, Sullivan R, Yusuf S (2015) The burden of disease in older people and implications for health policy and practice. Lancet 385(9967):549–562. https://doi.org/10.1016/S0140-6736(14)61347-7
Bruandet A, Richard F, Bombois S, Maurage CA, Deramecourt V, Lebert F et al (2009) Alzheimer disease with cerebrovascular disease and vascular dementia: clinical features and course compared with Alzheimer disease. J Neurol Neurosurg Psychiatry 80(2):133–139. https://doi.org/10.1136/jnnp.2007.137851
Cedazo-Minguez A, Winblad B (2010) Biomarkers for Alzheimer’s disease and other forms of dementia: clinical needs, limitations and future aspects. Exp Gerontol 45(1):5–14. https://doi.org/10.1016/j.exger.2009.09.008
Al-Qazzaz NK, Bin SH, Ali MD, Ahmad SA, Kalaivani Chellappan MD, Islam S, Escudero J (2014) Role of EEG as biomarker in the early detection and classification of dementia. Sci World J 2014:906038. https://doi.org/10.1155/2014/906038
Menon V, Crottaz-Herbette S (2005) Combined EEG and fMRI studies of human brain function. Int Rev Neurobiol 66:291–321. https://doi.org/10.1016/s0074-7742(05)66010-2
Nunez P, Srinivasan R (2006) Electric fields of the brain: the neurophysics of EEG. Oxford University Press
Dauwels J, Vialatte F, Musha T, Cichocki A (2010) A comparative study of synchrony measures for the early diagnosis of Alzheimer’s disease, based on EEG. Neuroimage 49(1):668–693. https://doi.org/10.1016/j.neuroimage.2009.06.056
Trambaiolli LR, Lorena AC, Fraga FJ, Kanda PAM, Renato Anghinah R, Nitrini, (2011) Improving Alzheimer’s disease diagnosis with machine learning techniques. Clin EEG Neurosci 42(3):160–165. https://doi.org/10.1177/155005941104200304
Podgorelec V (2012) Analyzing EEG signals with machine learning for diagnosing Alzheimer’s disease. Elektronika Ir Elektrotechnika 18(8):61–64. https://doi.org/10.5755/j01.eee.18.8.2627
Fiscon G et al (2014) Alzheimer’s disease patients classification through EEG signals processing. IEEE Symposium on Computational Intelligence and Data Mining (CIDM) 2014:105–112. https://doi.org/10.1109/CIDM.2014.7008655
Al-Nuaimi AH, Jammeh E, Sun L, Ifeachor E (2015) Tsallis entropy as a biomarker for detection of Alzheimer’s disease. Annu Int Conf IEEE Eng Med Biol Soc 2015:4166–4169. https://doi.org/10.1109/embc.2015.7319312
Claudio B, Triggiani Antonio I et al (2016) Classification of single normal and Alzheimer’s disease individuals from cortical sources of resting state EEG rhythms. Front Neurosci 10:47. https://doi.org/10.3389/fnins.2016.00047
Triggiani Antonio I, Vitoantonio B, Antonio B et al (2017) Classification of healthy subjects and Alzheimer’s disease patients with dementia from cortical sources of resting state EEG rhythms: a study using artificial neural networks. Front Neurosci 10:604. https://doi.org/10.3389/fnins.2016.00604
Blinowska KJ, Rakowski F, Kaminski M, De Vico FF, Del Percio C, Lizio R, Babiloni C (2017) Functional and effective brain connectivity for discrimination between Alzheimer’s patients and healthy individuals: a study on resting state EEG rhythms. Clin Neurophysiol 128(4):667–680. https://doi.org/10.1016/j.clinph.2016.10.002
Al-Nuaimi AH, Blūma M, Al-Juboori SS, Eke CS, Jammeh E, Sun L, Ifeachor E (2021) Robust EEG based biomarkers to detect Alzheimer’s disease. Brain Sci 11(8):1026. https://doi.org/10.3390/brainsci11081026
Bajaj V, Pachori RB (2012) Classification of seizure and non-seizure EEG signals using empirical mode decomposition. IEEE Trans Inf Technol Biomed 16(6):1135–1142. https://doi.org/10.1109/TITB.2011.2181403
Huang NE, Shen Z, Long SR et al (1998) The empirical mode decomposition and the Hilbert spectrum for nonlinear and nonstationary time series analysis. Proceedings of The Royal Society A Mathematical Physical and Engineering Sciences 454(1971):903–995. https://doi.org/10.1098/rspa.1998.0193
Cui D, Wang J, Bian Z, Li Q, Wang L, Li X (2015) Analysis of entropies based on empirical mode decomposition in amnesic mild cognitive impairment of diabetes mellitus. Journal of Innovative Optical Health 8(5):1550010. https://doi.org/10.1142/S1793545815500108
Lazar P, Jayapathy R, Jordina Torrents-Barrena M, Lindad M, Beena Mol J, Mohanalin DP (2018) Improving the performance of empirical mode decomposition via Tsallis entropy: application to Alzheimer EEG analysis. Biomed Mater Eng 29(5):551–566. https://doi.org/10.3233/bme-181008
Harati A, Lopez S, Obeid I, Jacobson M, Tobochnik S, Picone J (2014) THE TUH EEG CORPUS: a big data resource for automated EEG interpretation. IEEE Signal Processing in Medicine and Biology Symposium (SPMB) 2014:1–5. https://doi.org/10.1109/SPMB.2014.7002953
The Neural Engineering Data Consortium (NEDC), TUH EEG Corpus 2017. Available: https://www.isip.piconepress.com/projects/tuh_eeg/html/overview.shtml
Sean Ferrell, Vineetha Mathew, Matthew Refford, Vincent Tchiong, Tameem Ahsan, Iyad Obeid, Joseph Picone (2020) The Temple University Hospital EEG Corpus: electrode location and channel labels. In: Neural engineering data consortium report. Available: https://www.isip.piconepress.com/publications/reports/2020/tuh_eeg/electrodes/
Ho R, Hung K (2020) A comparative investigation of mode mixing in EEG decomposition using EMD, EEMD and M-EMD. IEEE Symposium on Computer Applications and Industrial Electronics (ISCAIE) 2020:203–210. https://doi.org/10.1109/ISCAIE47305.2020.9108817
Xua G, Yangb Z, Wang S (2016) Study on mode mixing problem of EMD. Joint International Information Technology, Mechanical and Electronic Engineering Conference 2016:389–394. https://doi.org/10.2991/jimec-16.2016.69
Zhu W, Zhao H, Xiang D, Chen X (2013) A flattest constrained envelope approach for empirical mode decomposition. PLoS ONE 8(4):e61739. https://doi.org/10.1371/journal.pone.0061739
Zhaohua Wu, Huang NE (2009) Ensemble empirical mode decomposition: a noise-assisted data analysis method. Adv Adapt Data Anal 1(1):1–41. https://doi.org/10.1142/S1793536909000047
Deering R, Kaiser JF (2005) The use of masking signal to improve empirical mode decomposition. Proceedings of IEEE International Conference on Acoustics, Speech Signal Processing 2005:485–488. https://doi.org/10.1109/ICASSP.2005.1416051
Chen Q, Huang N, Riemenschneider S et al (2006) A B-spline approach for empirical mode decompositions. Adv Comput Math 24:171–195. https://doi.org/10.1007/s10444-004-7614-3
Xuejun Z, Yan H, Dongsheng W (2020) Improved EMD based on piecewise cubic Hermite interpolation and mirror extension. Chin J Electron 29(5):899–905. https://doi.org/10.1049/cje.2020.08.005
Zhengguang Xu, Huang B, Li K (2010) An alternative envelope approach for empirical mode decomposition. Digit Signal Process 20(1):77–84. https://doi.org/10.1016/j.dsp.2009.06.009
Bejancu A (2000) A new approach to semi-cardinal spline interpolation. East J Approx 6(4):447–463
Andrzejak RG, Lehnertz K, Mormann F, Rieke C, David P, Elger CE (2001) Indications of nonlinear deterministic and finite-dimensional structures in time series of brain electrical activity: dependence on recording region and brain state. Phys Rev E Stat Nonlin Soft Matter Phys 64:061907. https://doi.org/10.1103/physreve.64.061907
Vecchio F, Babiloni C, Lizio R, De Vico FF, Blinowska K, Verrienti G, Frisoni G, Rossini PM (2013) Resting state cortical EEG rhythms in Alzheimer’s disease: toward EEG markers for clinical applications: a review. Suppl Clin Neurophysiol 62:223–236. https://doi.org/10.1016/b978-0-7020-5307-8.00015-6
Babiloni C et al (2015) Brain neural synchronization and functional coupling in Alzheimer’s disease as revealed by resting state EEG rhythms. Int J Psychophysiol 103:88–102. https://doi.org/10.1016/j.ijpsycho.2015.02.008
Vecchio F, Babiloni C (2011) Direction of information flow in Alzheimer’s disease and MCI patients. Int J Alzheimers Dis 2011:214580. 10.4061%2F2011%2F214580
Gallego-Jutglà E, Elgendi M, Vialatte F, Solé-Casals J, Cichocki A, Latchoumane C, Jeong J, Dauwels J (2012) Diagnosis of Alzheimer’s disease from EEG by means of synchrony measures in optimized frequency bands. Annu Int Conf IEEE Eng Med Biol Soc 2012:4266–4270. https://doi.org/10.1109/embc.2012.6346909
Seraj E (2018) Cerebral synchrony assessment: a general review on cerebral signals’ synchronization estimation concepts and methods. arXiv preprint arXiv:1612.04295. Available: https://doi.org/10.48550/arXiv.1612.04295
Faes L, Erla S, Nollo G (2012) Measuring connectivity in linear multivariate processes: definitions, interpretation, and practical analysis. Comput Math Methods Med 2012:140513. https://doi.org/10.1155/2012/140513
Locatelli T, Cursi M, Liberati D, Franceschi M, Comi G (1998) EEG coherence in Alzheimer’s disease. Electroencephalogr Clin Neurophysiol 106(3):229–237. https://doi.org/10.1016/S0013-4694(97)00129-6
Brunovsky M, Matousek M, Edman A, Cervena K, Krajca V (2003) Objective assessment of the degree of dementia by means of EEG. Neuropsychobiology 48(1):19–26. https://doi.org/10.1159/000071824
Combrisson E, Jerbi K (2015) Exceeding chance level by chance: the caveat of theoretical chance levels in brain signal classification and statistical assessment of decoding accuracy. J Neurosci Methods 250:126–136. https://doi.org/10.1016/j.jneumeth.2015.01.010
Beleites C, Baumgartner R, Bowman C, Somorjai RL, Steiner G, Salzer R, Sowa MG (2005) Variance reduction in estimating classification error using sparse datasets. Chemom Intell Lab Syst 79:91–100. https://doi.org/10.1016/J.CHEMOLAB.2005.04.008
Wainer J, Cawley GC (2017) Empirical evaluation of resampling procedures for optimising SVM hyperparameters. J Mach Learn Res 18(1):475−509. https://dl.acm.org/doi/abs/https://doi.org/10.5555/3122009.3122024
Janez Demšar (2006) Statistical comparisons of classifiers over multiple data sets. J Mach Learn Res 7:1–30. https://dl.acm.org/doi/https://doi.org/10.5555/1248547.1248548
Lehmann C, Koenig T, Jelic V, Prichep L, John RE, Wahlund LO, Dodge Y, Dierks T (2007) Application and comparison of classification algorithms for recognition of Alzheimer’s disease in electrical brain activity (EEG). J Neurosci Methods 161(2):342–350. https://doi.org/10.1016/j.jneumeth.2006.10.023
Riaz F, Hassan A, Rehman S, Niazi IK, Dremstrup K (2016) EMD-based temporal and spectral features for the classification of EEG signals using supervised learning. IEEE Trans Neural Syst Rehabil Eng 24(1):28–35. https://doi.org/10.1109/TNSRE.2015.2441835
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46:389–422. https://doi.org/10.1023/A:1012487302797
Byun H, Lee SW (2003) A survey on pattern recognition applications of support vector machines. Int J Pattern Recognit Artif Intell 17(3):459–486. https://doi.org/10.1142/S0218001403002460
Rilling G, Flandrin P, Gonalves P (2003) On empirical mode decomposition and its algorithms. IEEE-EURASIP Workshop on Nonlinear Signal and Image Processing NSIP-03. Codes Available: https://perso.ens-lyon.fr/patrick.flandrin/emd.html
Magrin-Chagnolleau I (2002) MATLAB code for EMD. Codes Available: http://www.mit.edu/~gari/CODE/HRV/emd.m
Khan M A, Yoshio Ohno (2007) An automated video data compression algorithm by cardinal spline fitting. NICOGRAPH International Conference.
Faes L, Erla S, Porta A, Nollo G (2013) A framework for assessing frequency domain causality in physiological time series with instantaneous effects. Philosophical Transactions A 371:20110618. https://doi.org/10.1098/rsta.2011.0618. Codes Available: http://www.lucafaes.net/emvar.html
BioSig for MATLAB toolbox, Codes Available: http://biosig.sourceforge.net/index.html
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ho, R., Hung, K. EEG analysis and classification based on cardinal spline empirical mode decomposition and synchrony features. Med Biol Eng Comput 60, 2359–2372 (2022). https://doi.org/10.1007/s11517-022-02615-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-022-02615-y