Abstract
Accurate segmentation of honeycomb lung lesions from lung CT images plays a crucial role in the diagnosis and treatment of various lung diseases. However, the availability of algorithms for automatic segmentation of honeycomb lung lesions remains limited. In this study, we propose a novel multi-scale cross-layer attention fusion network (MCAFNet) specifically designed for the segmentation of honeycomb lung lesions, taking into account their shape specificity and similarity to surrounding vascular shadows. The MCAFNet incorporates several key modules to enhance the segmentation performance. Firstly, a multiscale aggregation (MIA) module is introduced in the input part to preserve spatial information during downsampling. Secondly, a cross-layer attention fusion (CAF) module is proposed to capture multiscale features by integrating channel information and spatial information from different layers of the feature maps. Lastly, a bidirectional attention gate (BAG) module is constructed within the skip connection to enhance the model’s ability to filter out background information and focus on the segmentation target. Experimental results demonstrate the effectiveness of the proposed MCAFNet. On the honeycomb lung segmentation dataset, the network achieves an Intersection over Union (IoU) of 0.895, mean IoU (mIoU) of 0.921, and mean Dice coefficient (mDice) of 0.949, outperforming existing medical image segmentation algorithms. Furthermore, experiments conducted on additional datasets confirm the generalizability and robustness of the proposed model. The contribution of this study lies in the development of the MCAFNet, which addresses the lack of automated segmentation algorithms for honeycomb lung lesions. The proposed network demonstrates superior performance in accurately segmenting honeycomb lung lesions, thereby facilitating the diagnosis and treatment of lung diseases. This work contributes to the existing literature by presenting a novel approach that effectively combines multi-scale features and attention mechanisms for lung lesion segmentation. The code is available at https://github.com/Oran9er/MCAFNet.
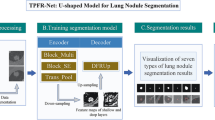
Graphical abstract















Similar content being viewed by others
References
Arakawa H, Honma K (2011) Honeycomb lung: history and current concepts. AJR Am J Roentgenol 196:773–782
Salisbury ML, Lynch DA, van Beek EJ, Kazerooni EA, Guo J, Xia M, Murray S, Anstrom KJ, Yow E, Martinez FJ, Hoffman EA, Flaherty KR, Investigators IP (2017) idiopathic pulmonary fibrosis: the association between the adaptive multiple features method and fibrosis outcomes. Am J Respir Crit Care Med 195:921–929
Bak SH, Park HY, Nam JH, Lee HY, Lee JH, Sohn I, Chung MP (2019) Predicting clinical outcome with phenotypic clusters using quantitative CT fibrosis and emphysema features in patients with idiopathic pulmonary fibrosis. PLoS One 14:e0215303
Norouzi A, Rahim MSM, Altameem A, Saba T, Rad AE, Rehman A, Uddin M (2014) Medical image segmentation methods, algorithms, and applications. IETE Tech Rev 31:199–213
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015, pp 234–241
Zhou Z, Siddiquee MMR, Tajbakhsh N, Liang J (2018) UNet++: a nested U-Net architecture for medical image segmentation. Deep Learn Med Image Anal Multimodal Learn Clin Decis Support 11045(2018):3–11
Alom MZ, Yakopcic C, Hasan M, Taha TM, Asari VK (2019) Recurrent residual U-Net for medical image segmentation. J Med Imaging (Bellingham) 6:014006
Chen J, Lu Y, Yu Q, Luo X, Adeli E, Wang Y, Lu L, Yuille AL, Zhou Y (2021) TransUNet: Transformers make strong encoders for medical image segmentation, pp arXiv:2102.04306
Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, Dehghani M, Minderer M, Heigold G, Gelly S (2020) An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929
Valanarasu JMJ, Patel VM (2022) UNeXt: MLP-based rapid medical image segmentation network. Medical Image Computing and Computer Assisted Intervention – MICCAI 2022, pp 23–33
Qureshi I, Yan J, Abbas Q, Shaheed K, Riaz AB, Wahid A, Khan MWJ, Szczuko P (2023) Medical image segmentation using deep semantic-based methods: a review of techniques, applications and emerging trends. Inf Fusion 90:316–352
Zhang H, Zhong X, Li G, Liu W, Liu J, Ji D, Li X, Wu J (2023) BCU-Net: bridging ConvNeXt and U-Net for medical image segmentation. Comput Biol Med 159:106960
Liu Z, Mao H, Wu C-Y, Feichtenhofer C, Darrell T, Xie S (2022) A ConvNet for the 2020s, pp arXiv:2201.03545
Qiu C, Liu Z, Song Y, Yin J, Han K, Zhu Y, Liu Y, Sheng VS (2023) RTUNet: residual transformer UNet specifically for pancreas segmentation. Biomed Signal Process Control 79:104173
Zhang X, Liu Y, Guo S, Song Z (2023) EG-Unet: edge-Guided cascaded networks for automated frontal brain segmentation in MR images. Comput Biol Med 158:106891
Chen L-C, Zhu Y, Papandreou G, Schroff F, Adam H (2018) Encoder-decoder with atrous separable convolution for semantic image segmentation. Proceedings of the European conference on computer vision (ECCV), pp 801–818
Sunnetci KM, Kaba E, Celiker FB, Alkan A (2023) Deep Network-Based Comprehensive Parotid Gland Tumor Detection. Acad Radiol. https://doi.org/10.1016/j.acra.2023.04.028
Jiang L, Ou J, Liu R, Zou Y, Xie T, Xiao H, Bai T (2023) RMAU-Net: residual multi-scale attention U-Net for liver and tumor segmentation in CT images. Comput Biol Med 158:106838
Chen L-C, Papandreou G, Kokkinos I, Murphy K, Yuille AL (2017) Deeplab: semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE Trans Pattern Anal Mach Intell 40:834–848
Kushnure DT, Talbar SN (2021) MS-UNet: a multi-scale UNet with feature recalibration approach for automatic liver and tumor segmentation in CT images. Comput Med Imaging Graph 89:101885
Wu R, Xin Y, Qian J, Dong Y (2023) A multi-scale interactive U-Net for pulmonary vessel segmentation method based on transfer learning. Biomed Signal Process Control 80:104407
Sun Y, Dai D, Zhang Q, Wang Y, Xu S, Lian C (2023) MSCA-Net: multi-scale contextual attention network for skin lesion segmentation. Patt Recog 139:109524
Xuan P, Jiang B, Cui H, Jin Q, Cheng P, Nakaguchi T, Zhang T, Li C, Ning Z, Guo M, Wang L (2022) Convolutional bi-directional learning and spatial enhanced attentions for lung tumor segmentation. Comput Methods Programs Biomed 226:107147
Fan D-P, Ji G-P, Zhou T, Chen G, Fu H, Shen J, Shao L (2020) Pranet: Parallel reverse attention network for polyp segmentation, Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, Peru, October 4–8, 2020, Proceedings, Part VI 23, Springer, pp 263–273
Zhang J, Qin Q, Ye Q, Ruan T (2023) ST-Unet: Swin Transformer boosted U-Net with cross-layer feature enhancement for medical image segmentation. Comput Biol Med 153:106516
Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, Lin S, Guo B (2021) Swin transformer: hierarchical vision transformer using shifted windows. pp arXiv:2103.14030
Ates GC, Mohan P, Celik E (2023) Dual cross-attention for medical image segmentation. pp arXiv:2303.17696
Chen L, Zhang H, Xiao J, Nie L, Shao J, Liu W, Chua T-S (2017) Sca-cnn: spatial and channel-wise attention in convolutional networks for image captioning. Proceedings of the IEEE conference on computer vision and pattern recognition. pp 5659–5667
Hu J, Shen L, Sun G (2018) Squeeze-and-excitation networks. Proceedings of the IEEE conference on computer vision and pattern recognition. pp 7132–7141
Oktay O, Schlemper J, Folgoc LL, Lee M, Heinrich M, Misawa K, Mori K, McDonagh S, Hammerla NY, Kainz B (2018) Attention u-net: learning where to look for the pancreas. arXiv preprint arXiv:1804.03999
Sun J, Zhang X, Li X, Liu R, Wang T (2023) DARMF-UNet: a dual-branch attention-guided refinement network with multi-scale features fusion U-Net for gland segmentation. Comput Biol Med 163:107218
Yu Q, Qi L, Gao Y, Wang W, Shi Y (2022) Crosslink-Net: double-branch encoder network via fusing vertical and horizontal convolutions for medical image segmentation. IEEE Trans Image Process 31:5893–5908
Qin C, Guerrero R, Bowles C, Chen L, Dickie DA, Valdes-Hernandez MdC, Wardlaw J, Rueckert D (2018) A large margin algorithm for automated segmentation of white matter hyperintensity. Pattern Recogn 77:150–159
Gao Z-K, Akben SB, Alkan A (2016) Visual interpretation of biomedical time series using Parzen window-based density-amplitude domain transformation. PloS One 11(9):e0163569
Han Z, Jian M, Wang G-G (2022) ConvUNeXt: an efficient convolution neural network for medical image segmentation. Knowl Based Syst 253:109512
Zhao X, Jia H, Pang Y, Lv L, Tian F, Zhang L, Sun W, Lu H (2023) M2SNet: multi-scale in multi-scale subtraction network for medical image segmentation. arXiv preprint arXiv:2303.10894
Jun M, Cheng G, Yixin W, Xingle A, Jiantao G, Ziqi Y, Minqing Z, Xin L, Xueyuan D, Shucheng C (2020) Covid-19 ct lung and infection segmentation dataset. 2020, Verson
Jha D, Smedsrud PH, Riegler MA, Halvorsen P, de Lange T, Johansen D, Johansen HD (2020) Kvasir-seg: a segmented polyp dataset, MultiMedia Modeling: 26th International Conference, MMM 2020, Daejeon, South Korea, January 5–8, 2020, Proceedings, Part II 26, Springer, pp 451-462
Bernal J, Sánchez FJ, Fernández-Esparrach G, Gil D, Rodríguez C, Vilariño F (2015) WM-DOVA maps for accurate polyp highlighting in colonoscopy: validation vs. saliency maps from physicians. Comput Med Imaging Graph 43:99–111
Vázquez D, Bernal J, Sánchez FJ, Fernández-Esparrach G, López AM, Romero A, Drozdzal M, Courville A (2017) A benchmark for endoluminal scene segmentation of colonoscopy images. J Healthc Eng 2017:1–9
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, G., Xie, J., Zhang, L. et al. MCAFNet: multiscale cross-layer attention fusion network for honeycomb lung lesion segmentation. Med Biol Eng Comput 62, 1121–1137 (2024). https://doi.org/10.1007/s11517-023-02995-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02995-9