Abstract
Purpose
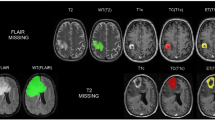
The gene mutation status of isocitrate dehydrogenase (IDH) in gliomas leads to a different prognosis. It is challenging to perform automated tumor segmentation and genotype prediction directly using label-deprived multimodal magnetic resonance (MR) images. We propose a novel framework that employs a domain adaptive mechanism to address this issue.
Methods
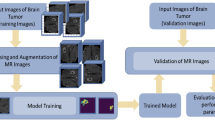
Multimodal domain adaptive segmentation (MDAS) framework was proposed to solve the gap issue in cross dataset model transfer. Image translation was used to adaptively align the multimodal data from two domains at the image level, and segmentation consistency loss was proposed to retain more pathological information through semantic constraints. The data distribution between the labeled public dataset and label-free target dataset was learned to achieve better unsupervised segmentation results on the target dataset. Then, the segmented tumor foci were used as a mask to extract the radiomics and deep features. And the subsequent prediction of IDH gene mutation status was conducted by training a random forest classifier. The prediction model does not need any expert segmented labels.
Results
We implemented our method on the public BraTS 2019 dataset and 110 astrocytoma cases of grade II–IV brain tumors from our hospital. We obtained a Dice score of 77.41% for unsupervised tumor segmentation, a genotype prediction accuracy (ACC) of 0.7639 and an area under curve (AUC) of 0.8600. Experimental results demonstrate that our domain adaptive approach outperforms the methods utilizing direct transfer learning. The model using hybrid features gives better results than the model using radiomics or deep features alone.
Conclusions
Domain adaptation enables the segmentation network to achieve better performance, and the extraction of mixed features at multiple levels on the segmented region of interest ensures effective prediction of the IDH gene mutation status.



Similar content being viewed by others
References
Parsons DW, Jones S, Zhang X, Lin JCH, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu I-M, Gallia GL (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321(5897):1807–1812. https://doi.org/10.1126/science.1164382
Oronsky B, Reid TR, Oronsky A, Sandhu N, Knox SJ (2021) A review of newly diagnosed glioblastoma. Front Oncol 10:574012. https://doi.org/10.3389/fonc.2020.574012
Louis DN, Perry A, Reifenberger G, Von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Akintola O, Samore W, Martinez-Lage Alvarez M, Gerstner ER (2020) Clinical, radiologic & prognostic profile of IDH wild type diffuse astrocytic glioma with molecular features of glioblastoma. J Clinic Oncol 38(15_suppl):2557. https://doi.org/10.1200/JCO.2020.38.15_suppl.2557
Preusser M, Wöhrer A, Stary S, Höftberger R, Streubel B, Hainfellner JA (2011) Value and limitations of immunohistochemistry and gene sequencing for detection of the IDH1-R132H mutation in diffuse glioma biopsy specimens. J Neuropathol Exp Neurol 70(8):715–723. https://doi.org/10.1097/nen.0b013e31822713f0
Smits M, van den Bent MJ (2017) Imaging correlates of adult glioma genotypes. Radiology 284(2):316–331. https://doi.org/10.1148/radiol.2017151930
Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby J, Burren Y, Porz N, Slotboom J, Wiest R (2014) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imaging 34(10):1993–2024. https://doi.org/10.1109/TMI.2014.2377694
van Opbroek A, Ikram MA, Vernooij MW (2014) Transfer learning improves supervised image segmentation across imaging protocols. IEEE Trans Med Imaging 34(5):1018–1030
Perone CS, Ballester P, Barros RC, Cohen-Adad J (2019) Unsupervised domain adaptation for medical imaging segmentation with self-ensembling. Neuroimage 194:1–11. https://doi.org/10.1016/j.neuroimage.2019.03.026
Torralba A, Efros AA (2011) Unbiased look at dataset bias. In: IEEE conference on computer vision and pattern recognition (CVPR), pp 1521–1528. https://doi.org/10.1109/CVPR.2011.5995347
Bousmalis K, Silberman N, Dohan D, Erhan D, Krishnan D (2017) Unsupervised pixel-level domain adaptation with generative adversarial networks. In: IEEE conference on computer vision and pattern recognition (CVPR), pp 3722–3731. https://doi.org/10.1109/CVPR.2017.18
Zhang Y, Miao S, Mansi T, Liao R (2018) Task driven generative modeling for unsupervised domain adaptation: application to x-ray image segmentation. In: International conference on medical image computing and computer-assisted intervention (MICCAI), pp 599–607. https://doi.org/10.1007/978-3-030-00934-2_67
Zhu J-Y, Park T, Isola P, Efros AA (2017) Unpaired image-to-image translation using cycle-consistent adversarial networks. In: Proceedings of the IEEE international conference on computer vision (ICCV), pp 2223–2232. https://doi.org/10.1109/ICCV.2017.244
Jiang J, Hu Y-C, Tyagi N, Zhang P, Rimner A, Mageras GS, Deasy JO, Veeraraghavan H (2018) Tumor-aware, adversarial domain adaptation from CT to MRI for lung cancer segmentation. In: International conference on medical image computing and computer-assisted intervention (MICCAI), pp 777–785. https://doi.org/10.1007/978-3-030-00934-2_86
Zhang Z, Yang L, Zheng Y (2018) Translating and segmenting multimodal medical volumes with cycle-and shape-consistency generative adversarial network. In: Proceedings of the IEEE conference on computer vision and pattern recognition (CVPR), pp 9242–9251.
Kamnitsas K, Baumgartner C, Ledig C, Newcombe V, Simpson J, Kane A, Menon D, Nori A, Criminisi A, Rueckert D (2017) Unsupervised domain adaptation in brain lesion segmentation with adversarial networks. In: International conference on information processing in medical imaging (IPMI), pp 597–609.
Chen C, Dou Q, Chen H, Qin J, Heng PA (2020) Unsupervised bidirectional cross-modality adaptation via deeply synergistic image and feature alignment for medical image segmentation. IEEE Trans Med Imaging 39(7):2494–2505. https://doi.org/10.1109/TMI.2020.2972701
Lu C-F, Hsu F-T, Hsieh KL-C, Kao Y-CJ, Cheng S-J, Hsu JB-K, Tsai P-H, Chen R-J, Huang C-C, Yen Y (2018) Machine learning–based radiomics for molecular subtyping of gliomas. Clin Cancer Res 24(18):4429–4436. https://doi.org/10.1158/1078-0432.CCR-17-3445
Lambin P, Leijenaar RT, Deist TM, Peerlings J, De Jong EE, Van Timmeren J, Sanduleanu S, Larue RT, Even AJ, Jochems A (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14(12):749–762. https://doi.org/10.1038/nrclinonc.2017.141
Pasquini L, Napolitano A, Tagliente E, Dellepiane F, Lucignani M, Vidiri A, Ranazzi G, Stoppacciaro A, Moltoni G, Nicolai M (2021) Deep learning can differentiate IDH-mutant from IDH-wild GBM. J Personalized Med 11(4):290. https://doi.org/10.3390/jpm11040290
Buda M, Saha A, Mazurowski MA (2019) Association of genomic subtypes of lower-grade gliomas with shape features automatically extracted by a deep learning algorithm. Comput Biol Med 109:218–225
Bangalore Yogananda CG, Shah BR, Vejdani-Jahromi M, Nalawade SS, Murugesan GK, Yu FF, Pinho MC, Wagner BC, Mickey B, Patel TR (2020) A novel fully automated MRI-based deep-learning method for classification of IDH mutation status in brain gliomas. Neuro Oncol 22(3):402–411. https://doi.org/10.1101/757385
Choi YS, Bae S, Chang JH, Kang S-G, Kim SH, Kim J, Rim TH, Choi SH, Jain R, Lee S-K (2021) Fully automated hybrid approach to predict the IDH mutation status of gliomas via deep learning and radiomics. Neuro Oncol 23(2):304–313. https://doi.org/10.1093/neuonc/noaa177
van Griethuysen JJM, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RGH, Fillon-Robin JC, Pieper S, Aerts HJWL (2017) Computational radiomics system to decode the radiographic phenotype. Can Res 77(21):e104–e107. https://doi.org/10.1158/0008-5472.CAN-17-0339
Zwanenburg A, Vallières M, Abdalah MA (2020) The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2):328–338
Simonyan K, Zisserman A (2015) Very deep convolutional networks for large-scale image recognition. In: International conference on learning representations (ICLR), pp 1–14.
Deng J, Dong W, Socher R, Li L-J, Li K, Fei-Fei L (2009) Imagenet: a large-scale hierarchical image database. In: 2009 IEEE conference on computer vision and pattern recognition (CVPR), pp 248–255. https://doi.org/10.1109/CVPR.2009.5206848
Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH (2021) nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 18(2):203–211
Huang H, Lin L, Tong R, Hu H, Zhang Q, Iwamoto Y, Han X, Chen Y-W, Wu J (2020) UNet 3+: A full-scale connected UNet for medical image segmentation. In: ICASSP 2020−2020 IEEE international conference on acoustics, speech and signal processing (ICASSP), pp 1055–1059. https://doi.org/10.1109/ICASSP40776.2020.9053405
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China under grants 82071913 and 82071869, in part by the Joint Scientific Research Foundation of Fujian Provincial Education and Health Commission of China under grant 2019-WJ-10, and in part by Science and Technology Project of Fujian Province under grant 2019Y0001.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards
Informed consent
As a retrospective study, the individual informed consent was waived.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zeng, H., Xing, Z., Gao, F. et al. A multimodal domain adaptive segmentation framework for IDH genotype prediction. Int J CARS 17, 1923–1931 (2022). https://doi.org/10.1007/s11548-022-02700-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-022-02700-5