Abstract
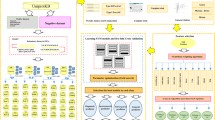
Fertility is the most crucial step in the development process, which is controlled by many fertility-related proteins, including spermatogenesis-, oogenesis- and embryogenesis-related proteins. The identification of fertility-related proteins can provide important clues for studying the role of these proteins in development. Therefore, in this study, we constructed a two-layer classifier to identify fertility-related proteins. In this classifier, we first used the composition of amino acids (AA) and their physical and chemical properties to code these three fertility-related proteins. Then, the feature set is optimized by analysis of variance (ANOVA) and incremental feature selection (IFS) to obtain the optimal feature subset. Through five-fold cross-validation (CV) and independent data tests, the performance of models constructed by different machine learning (ML) methods is evaluated and compared. Finally, based on support vector machine (SVM), we obtained a two-layer model to classify three fertility-related proteins. On the independent test data set, the accuracy (ACC) and the area under the receiver operating characteristic curve (AUC) of the first layer classifier are 81.95% and 0.89, respectively, and them of the second layer classifier are 84.74% and 0.90, respectively. These results show that the proposed model has stable performance and satisfactory prediction accuracy, and can become a powerful model to identify more fertility related proteins.
Similar content being viewed by others
Explore related subjects
Discover the latest articles and news from researchers in related subjects, suggested using machine learning.References
Johnson J, Bagley J, Skaznik-Wikiel M, Lee H J, Adams G B, Niikura Y, Tschudy K S, Tilly J C, Cortes M L, Forkert R, Spitzer T, Iacomini J, Scadden D T, Tilly J L. Oocyte generation in adult mammalian ovaries by putative germ cells in bone marrow and peripheral blood. Cell, 2005, 122(2): 303–315
Neto F T L, Bach P V, Najari B B, Li P S, Goldstein M. Spermatogenesis in humans and its affecting factors. Seminars in Cell & Developmental Biology, 2016, 59: 10–26
Müller F, Tora L. TBP2 is a general transcription factor specialized for female germ cells. Journal of Biology, 2009, 8(11): 97
Izaguirre M F, Casco V H. E-cadherin roles in animal biology: a perspective on thyroid hormone-influence. Cell Communication and Signaling, 2016, 14(1): 27
Rahimi M, Bakhtiarizadeh M R, Mohammadi-Sangcheshmeh A. OOgenesis_Pred: a sequence-based method for predicting oogenesis proteins by six different modes of Chou’s pseudo amino acid composition. Journal of Theoretical Biology, 2017, 414: 128–136
Bakhtiarizadeh M R, Rahimi M, Mohammadi-Sangcheshmeh A, Shariati J V, Salami S A. PrESOgenesis: a two-layer multi-label predictor for identifying fertility-related proteins using support vector machine and pseudo amino acid composition approach. Scientific Reports, 2018, 8(1): 9025
Le N Q K. Fertility-GRU: Identifying fertility-related proteins by incorporating deep-gated recurrent units and original position-specific scoring matrix profiles. Journal of Proteome Research, 2019, 18(9): 3503–3511
Wu X, Yu L. EPSOL: sequence-based protein solubility prediction using multidimensional embedding. Bioinformatics, 2021, 37(23): 4314–4320
Liu Q, Wan J, Wang G. A survey on computational methods in discovering protein inhibitors of SARS-CoV-2. Briefings in Bioinformatics, 2022, 23(1): bbab416
Zhao X, Wang H, Li H, Wu Y, Wang G. Identifying plant pentatricopeptide repeat proteins using a variable selection method. Frontiers in Plant Science, 2021, 12: 506681
Tao Z, Li Y, Teng Z, Zhao Y. A method for identifying vesicle transport proteins based on LibSVM and MRMD. Computational and Mathematical Methods in Medicine, 2020, 2020: 8926750
Guo Z, Wang P, Liu Z, Zhao Y. Discrimination of thermophilic proteins and non-thermophilic proteins using feature dimension reduction. Frontiers in Bioengineering and Biotechnology, 2020, 8: 584807
Zhang Q, Li H, Liu Y, Li J, Wu C, Tang H. Exosomal non-coding RNAs: new insights into the biology of hepatocellular carcinoma. Current Oncology, 2022, 29(8): 5383–5406
The UniProt Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Research, 2021, 49(D1): D480–D489
Hasan M M, Tsukiyama S, Cho J Y, Kurata H, Alam A, Liu X, Manavalan B, Deng H W. Deepm5C: a deep-learning-based hybrid framework for identifying human RNA N5-methylcytosine sites using a stacking strategy. Molecular Therapy, 2022, 30(8): 2856–2867
Jeon Y J, Hasan M, Park H W, Lee K W, Manavalan B. TACOS: a novel approach for accurate prediction of cell-specific long noncoding RNAs subcellular localization. Briefings in Bioinformatics, 2022, 23(4): bbac243
Chen Z, Zhao P, Li F, Leier A, Marquez-Lago T T, Wang Y, Webb G I, Smith A I, Daly R J, Chou K C, Song J. iFeature: a Python package and Web server for features extraction and selection from protein and peptide sequences. Bioinformatics, 2018, 34(14): 2499–2502
Awais M, Hussain W, Rasool N, Khan Y D. iTSP-PseAAC: identifying tumor suppressor proteins by using fully connected neural network and PseAAC. Current Bioinformatics, 2021, 16(5): 700–709
Romdhane T F, Alhichri H, Ouni R, Atri M. Electrocardiogram heartbeat classification based on a deep convolutional neural network and focal loss. Computers in Biology and Medicine, 2020, 123: 103866
Alguwaizani S, Ren S, Huang D S, Han K. Predicting interactions between pathogen and human proteins based on the relation between sequence length and amino acid composition. Current Bioinformatics, 2021, 16(6): 799–806
Yu L, Wang M, Yang Y, Xu F, Zhang X, Xie F, Gao L, Li X. Predicting therapeutic drugs for hepatocellular carcinoma based on tissue-specific pathways. PLoS Computational Biology, 2021, 17(2): e1008696
Ahmed Z, Zulfiqar H, Khan A A, Gul I, Dao F Y, Zhang Z Y, Yu X L, Tang L. iThermo: a sequence-based model for identifying thermophilic proteins using a multi-feature fusion strategy. Frontiers in Microbiology, 2022, 13: 790063
Bian H, Guo M, Wang J. Recognition of mitochondrial proteins in plasmodium based on the tripeptide composition. Frontiers in Cell and Developmental Biology, 2020, 8: 578901
Hosen F, Mahmud S M H, Ahmed K, Chen W, Moni M A, Deng H W, Shoombuatong W, Hasan M. DeepDNAbP: a deep learning-based hybrid approach to improve the identification of deoxyribonucleic acid-binding proteins. Computers in Biology and Medicine, 2022, 145: 105433
Yang L, Gao H, Wu K, Zhang H, Li C, Tang L. Identification of cancerlectins by using cascade linear discriminant analysis and optimal g-gap tripeptide composition. Current Bioinformatics, 2020, 15(6): 528–537
Feng Z P, Zhang C T. Prediction of membrane protein types based on the hydrophobic index of amino acids. Journal of Protein Chemistry, 2000, 19(4): 269–275
Sokal R R, Thomson B A. Population structure inferred by local spatial autocorrelation: an example from an Amerindian tribal population. American Journal of Physical Anthropology, 2006, 129(1): 121–131
Horne D S. Prediction of protein helix content from an autocorrelation analysis of sequence hydrophobicities. Biopolymers, 1988, 27(3): 451–477
Hasan M, Schaduangrat N, Basith S, Lee G, Shoombuatong W, Manavalan B. HLPpred-Fuse: improved and robust prediction of hemolytic peptide and its activity by fusing multiple feature representation. Bioinformatics, 2020, 36(11): 3350–3356
Manavalan B, Patra M C. MLCPP 2.0: an updated cell-penetrating peptides and their uptake efficiency predictor. Journal of Molecular Biology, 2022, 434(11): 167604
Wang J, Zhang L, Jia L, Ren Y, Yu G. Protein-protein interactions prediction using a novel local conjoint triad descriptor of amino acid sequences. International Journal of Molecular Sciences, 2017, 18(11): 2373
Chou K C. Prediction of protein cellular attributes using pseudo-amino acid composition. Proteins, 2001, 43(3): 246–255
Naseer S, Hussain W, Khan Y D, Rasool N. NPalmitoylDeep-pseAAC: a predictor of N-palmitoylation sites in proteins using deep representations of proteins and PseAAC via modified 5-steps rule. Current Bioinformatics, 2021, 16(2): 294–305
Lv H, Yan K, Guo Y, Zou Q, Hesham A E L, Liu B. AMPpred-EL: an effective antimicrobial peptide prediction model based on ensemble learning. Computers in Biology and Medicine, 2022, 146: 105577
Chou K C. Using amphiphilic pseudo amino acid composition to predict enzyme subfamily classes. Bioinformatics, 2005, 21(1): 10–19
Dao F Y, Lv H, Zhang Z Y, Lin H. BDselect: a package for k-mer selection based on the binomial distribution. Current Bioinformatics, 2022, 17(3): 238–244
Shaban T F, Alkawareek M Y. Prediction of qualitative antibiofilm activity of antibiotics using supervised machine learning techniques. Computers in Biology and Medicine, 2022, 140: 105065
Ao C, Zou Q, Yu L. RFhy-m2G: identification of RNA N2-methylguanosine modification sites based on random forest and hybrid features. Methods, 2021, 203: 32–39
Gao S, Wang P, Feng Y, Xie X, Duan M, Fan Y, Liu S, Huang L, Zhou F. RIFS2D: a two-dimensional version of a randomly restarted incremental feature selection algorithm with an application for detecting low-ranked biomarkers. Computers in Biology and Medicine, 2021, 133: 104405
Rigatti S J. Random forest. Journal of Insurance Medicine, 2017, 47(1): 31–39
Ao C, Zou Q, Yu L. NmRF: identification of multispecies RNA 2’-O-methylation modification sites from RNA sequences. Briefings in Bioinformatics, 2021, 23(1): bbab480
Nakayama J Y, Ho J, Cartwright E, Simpson R, Hertzberg V S. Predictors of progression through the cascade of care to a cure for hepatitis C patients using decision trees and random forests. Computers in Biology and Medicine, 2021, 134: 104461
Jog A, Carass A, Roy S, Pham D L, Prince J L. Random forest regression for magnetic resonance image synthesis. Medical Image Analysis, 2017, 35: 475–488
Wu C, Lin B, Shi K, Zhang Q, Gao R, Yu Z, De Marinis Y, Zhang Y, Liu Z P. PEPRF: identification of essential proteins by integrating topological features of PPI network and sequence-based features via random forest. Current Bioinformatics, 2021, 16(9): 1161–1168
Huang Y, Zhou D, Wang Y, Zhang X, Su M, Wang C, Sun Z, Jiang Q, Sun B, Zhang Y. Prediction of transcription factors binding events based on epigenetic modifications in different human cells. Epigenomics, 2020, 12(16): 1443–1456
Basith S, Lee G, Manavalan B. STALLION: a stacking-based ensemble learning framework for prokaryotic lysine acetylation site prediction. Briefings in Bioinformatics, 2022, 23(1): bbab376
Shoombuatong W, Basith S, Pitti T, Lee G, Manavalan B. THRONE: a new approach for accurate prediction of human RNA N7-methylguanosine sites. Journal of Molecular Biology, 2022, 434(11): 167549
Cui Y, Zhai Y L, Qi Y Y, Liu X R, Zhao Y F, Lv F, Han L P, Zhao Z Z. The comprehensive analysis of clinical trials registration for IgA nephropathy therapy on ClinicalTrials. gov. Renal Failure, 2022, 44(1): 461–472
Chen C, Shi H, Jiang Z, Salhi A, Chen R, Cui X, Yu B. DNN-DTIs: improved drug-target interactions prediction using XGBoost feature selection and deep neural network. Computers in Biology and Medicine, 2021, 136: 104676
Hutchinson N, Klas K, Carlisle B G, Kimmelman J, Waligora M. How informative were early SARS-CoV-2 treatment and prevention trials? A longitudinal cohort analysis of trials registered on ClinicalTrials gov. PLoS One, 2022, 17(1): e0262114
Yang H, Luo Y, Ren X, Wu M, He X, Peng B, Deng K, Yan D, Tang H, Lin H. Risk prediction of diabetes: big data mining with fusion of multifarious physical examination indicators. Information Fusion, 2021, 75: 140–149
Dao F Y, Lv H, Zulfiqar H, Yang H, Su W, Gao H, Ding H, Lin H. A computational platform to identify origins of replication sites in eukaryotes. Briefings in Bioinformatics, 2021, 22(2): 1940–1950
Joshi P, Vedhanayagam M, Ramesh R. An ensembled SVM based approach for predicting adverse drug reactions. Current Bioinformatics, 2021, 16(3): 422–432
Usman S M, Khalid S, Bashir S. A deep learning based ensemble learning method for epileptic seizure prediction. Computers in Biology and Medicine, 2021, 136: 104710
Jiang Q, Wang G, Jin S, Li Y, Wang Y. Predicting human microRNA-disease associations based on support vector machine. International Journal of Data Mining and Bioinformatics, 2013, 8(3): 282–293
Yu L, Xia M, An Q. A network embedding framework based on integrating multiplex network for drug combination prediction. Briefings in Bioinformatics, 2021, 23(1): bbab364
Zhang S, Jiang H, Gao B, Yang W, Wang G. Identification of diagnostic markers for breast cancer based on differential gene expression and pathway network. Frontiers in Cell and Developmental Biology, 2022, 9: 811585
Sun Z, Huang Q, Yang Y, Li S, Lv H, Zhang Y, Lin H, Ning L. PSnoD: identifying potential snoRNA-disease associations based on bounded nuclear norm regularization. Briefings in Bioinformatics, 2022, 23(4): bbac240
Xu Z, Luo M, Lin W, Xue G, Wang P, Jin X, Xu C, Zhou W, Cai Y, Yang W, Nie H, Jiang Q. DLpTCR: an ensemble deep learning framework for predicting immunogenic peptide recognized by T cell receptor. Briefings in Bioinformatics, 2021, 22(6): bbab335
Lv Z, Wang P, Zou Q, Jiang Q. Identification of sub-Golgi protein localization by use of deep representation learning features. Bioinformatics, 2020, 36(24): 5600–5609
Song G, Wang G, Luo X, Cheng Y, Song Q, Wan J, Moore C, Song H, Jin P, Qian J, Zhu H. An all-to-all approach to the identification of sequence-specific readers for epigenetic DNA modifications on cytosine. Nature Communications, 2021, 12(1): 795
Lv H, Dao F Y, Lin H. DeepKla: an attention mechanism-based deep neural network for protein lysine lactylation site prediction. iMeta, 2022, 1(1): e11
Kopylov A T, Papysheva O, Gribova I, Kaysheva A L, Kotaysch G, Kharitonova L, Mayatskaya T, Nurbekov M K, Schipkova E, Terekhina O, Morozov S G. Severe types of fetopathy are associated with changes in the serological proteome of diabetic mothers. Medicine, 2021, 100(45): e27829
Pla I, Sanchez A, Pors S E, Pawlowski K, Appelqvist R, Sahlin K B, La Cour Poulsen L, Marko-Varga G, Andersen C Y, Malm J. Proteome of fluid from human ovarian small antral follicles reveals insights in folliculogenesis and oocyte maturation. Human Reproduction, 2021, 36(3): 756–770
Li C, Song C, Qi K, Liu Y, Dou Y, Li X, Qiao R, Wang K, Han X, Li X. Identification of estrus in sows based on salivary proteomics. Animals, 2022, 12(13): 1656
Li D Y, Yang X X, Tu C F, Wang W L, Meng L L, Lu G X, Tan Y Q, Zhang Q J, Du J. Sperm flagellar 2 (SPEF2) is essential for sperm flagellar assembly in humans. Asian Journal of Andrology, 2022, 24(4): 359–366
Zhang Z Y, Ning L, Ye X, Yang Y H, Futamura Y, Sakurai T, Lin H. iLoc-miRNA: extracellular/intracellular miRNA prediction using deep BiLSTM with attention mechanism. Briefings in Bioinformatics, 2022, 23(5): bbac395
Zhang L, Yang Y, Chai L, Li Q, Liu J, Lin H, Liu L. A deep learning model to identify gene expression level using cobinding transcription factor signals. Briefings in Bioinformatics, 2022, 23(1): bbab501
Acknowledgements
This research was funded by the Sichuan Major Science and Technology Project (2021ZDZX0009); the National Natural Science Foundation of China (Grant No. 035Z2060).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yan Lin, a professor of Animal Nutrition Institute at Sichuan Agricultural University, China. Her research is in the areas of animal nutrition and feed science.
Jiashu Wang, a master candidate of Center for Informational Biology, School of Life Science and Technology, University of Electronic Science and Technology of China, China. His research interests include bioinformatics and machine learning.
Xiaowei Liu, a master candidate of Center for Informational Biology, School of Life Science and Technology, University of Electronic Science and Technology of China, China. Her research interests include bioinformatics, machine learning and drug development.
Xueqin Xie, a master candidate of Center for Informational Biology, School of Life Science and Technology, University of Electronic Science and Technology of China, China. Her research interests are bioinformatics, machine learning and biomarker prediction.
De Wu, a professor of Animal Nutrition Institute at Sichuan Agricultural University, China. His research is in the areas of animal nutrition and feed science.
Junjie Zhang, a professor of College of Life Science at Sichuan Agricultural University, China. His research is in the areas of molecular biology of starch synthesis in maize.
Hui Ding, an associate professor of Center for Informational Biology at University of Electronic Science and Technology of China, China. Her research is in the areas of computational biology and system biology.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Lin, Y., Wang, J., Liu, X. et al. A computational model to identify fertility-related proteins using sequence information. Front. Comput. Sci. 18, 181902 (2024). https://doi.org/10.1007/s11704-022-2559-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11704-022-2559-6