Abstract
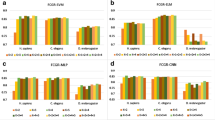
Nucleosome positioning played significant roles in various biological processes. With the development of high-throughput techniques, many methods and software were developed for nucleosome positioning. Although results with high accuracy (Acc) were obtained, the key factors for determining nucleosome positioning under less time complexity remain unresolved. Therefore, combining generalized relative entropy with self-similarity of DNA sequences, a novel method of nucleosome positioning was proposed for predicting nucleosome positioning in human, worm, fly and yeast genomes, respectively. Experimental results showed that prediction Acc of nucleosome positioning in aforementioned datasets reached 87.78%, 87.98%, 83.36% and 100%, respectively. Furthermore, it was found that five-nucleotide and six-nucleotide sequences were the determinant factors in nucleosome positioning.











Similar content being viewed by others
References
Astrovskaya I, Tork B, Mangul S, Westbrooks K, Mandoiu I, Balfe P, Zelikovsky A (2011) Inferring viral quasispecies spectra from 454 pyrosequencing reads. BMC Bioinform 12(Suppl6):S1. https://doi.org/10.1186/1471-2105-12-S6-S1
Awazu A (2017) Prediction of nucleosome positioning by the incorporation of frequencies and distributions of three different nucleotide segment lengths into a general pseudo k-tuple nucleotide composition. Bioinformatics 33(1):42–48. https://doi.org/10.1093/bioinformatics/btw562
Beigi S, Gohari A (2014) Quantum achievability proof via collision relative entropy. IEEE Trans Inf Theory 60(12):7980–7986. https://doi.org/10.1109/TIT.2014.2361632
Benson G (2002) A new distance measure for comparing sequence profiles based on path lengths along an entropy surface. Bioinformatics 18(suppl_2):S44–S53. https://doi.org/10.1093/bioinformatics/18.suppl_2.s44
Berbenetz NM, Nislow C, Brown GW (2010) Diversity of eukaryotic DNA replication origins revealed by genome-wide analysis of chromatin structure. PLoS Genet. https://doi.org/10.1371/journal.pgen.1001092
Bhasin M, Raghava G (2004) ESLpred: SVM-based method for subcellular localization of eukaryotic proteins using dipeptide composition and PSI-BLAST. Nucl Acids Res 32(suppl_2):W414–W419. https://doi.org/10.1093/nar/gkh350
Chen H, Zhou L (2012) A relative entropy approach to group decision making with interval reciprocal relations based on COWA operator. Group Decis Negot 21(4):585–599. https://doi.org/10.1007/s10726-011-9228-8
Chen W, Lin H, Feng PM, Ding C, Zuo YC, Chou KC (2012) iNuc-PhysChem: a sequence-based predictor for identifying nucleosomes via physicochemical properties. PLoS ONE 7(10):e47843. https://doi.org/10.1371/journal.pone.0047843
Chen W, Lin H, Chou KC (2015) Pseudo nucleotide composition or PseKNC: an effective formulation for analyzing genomic sequences. Mol BioSyst 11(10):2620–2634. https://doi.org/10.1039/C5MB00155B
Chen W, Feng P, Ding H, Lin H, Chou KC (2016) Using deformation energy to analyze nucleosome positioning in genomes. Genomics 107(2–3):69–75. https://doi.org/10.1016/j.ygeno.2015.12.005
Fabris F, Doherty A, Palmer D, de Magalhaes JP, Freitas AA (2018) A new approach for interpreting random forest models and its application to the biology of ageing. Bioinformatics 34(14):2449–2456. https://doi.org/10.1093/bioinformatics/bty087
Flores O, Orozco M (2011) nucleR: a package for nonparametric nucleosome positioning. Bioinformatics 27(15):2149–2150. https://doi.org/10.1093/bioinformatics/btr345
Freeman GS, Lequieu JP, Hinckley DM, de Pablo J (2014) DNA shape dominates sequence affinity in nucleosome formation. Phys Rev Lett 113(16):168101. https://doi.org/10.1103/PhysRevLett.113.168101
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28(23):3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Gibb S, Strimmer K (2015) Differential protein expression and peak selection in mass spectrometry data by binary discriminant analysis. Bioinformatics 31(19):3156–3162. https://doi.org/10.1093/bioinformatics/btv334
Guo SH, Deng EZ, Xu LQ, Ding H, Lin H, Chen W, Chou KC (2014) iNuc-PseKNC: a sequence-based predictor for predicting nucleosome positioning in genomes with pseudo k-tuple nucleotide composition. Bioinformatics 30(11):1522–1529. https://doi.org/10.1093/bioinformatics/btu083
Ide H, Umezawa M, Ohwada H (2016) Function prediction of disease-related long intergenic non-coding rna using random forest. In: Proceedings of the 7th international conference on computational systems-biology and bioinformatics. https://doi.org/10.1145/3029375.3029384
Ioshikhes I, Bolshoy A, Derenshteyn K, Borodovsky M, Trifonov EN (1996) Nucleosome dna sequence pattern revealed by multiple alignment of experimentally mapped sequences. J Mol Biol 262(2):129–139. https://doi.org/10.1006/jmbi.1996.0503
Ioshikhes IP, Albert I, Zanton SJ, Pugh BF (2006) Nucleosome positions predicted through comparative genomics. Nat Genet 38(10):1210–1215. https://doi.org/10.1038/ng1878
Ismail H, Saigo H, Dukka K (2017) RF-NR: random forest based approach for improved classification of nuclear receptors. IEEE/ACM Trans Comput Biol Bioinform. https://doi.org/10.1109/tcbb.2017.2773063
Kaplan N, Moore IK, Fondufe-Mittendorf Y, Gossett AJ, Tillo D, Field Y, LeProust EM, Hughes TR, Lieb JD, Widom J et al (2009) The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458:362–366. https://doi.org/10.1038/nature07667
Karlekar NP, Gomathi N (2018) OW-SVM: ontology and whale optimization-based support vector machine for privacy-preserved medical data classification in cloud. Int J Commun Syst. https://doi.org/10.1002/dac.3700
Lee W, Tillo D, Bray N, Morse RH, Davis RW, Hughes TR, Nislow C (2007) A high-resolution atlas of nucleosome occupancy in yeast. Nat Genet 39:1235–1244. https://doi.org/10.1038/ng2117
Lin W, Ji D, Lu Y (2017) Disorder recognition in clinical texts using multi-label structured SVM. BMC Bioinform 18:75. https://doi.org/10.1186/s12859-017-1476-4
Liu H, Duan X, Yu S, Sun X (2011) Analysis of nucleosome positioning determined by DNA helix curvature in the human genome. BMC Genomics 12:72. https://doi.org/10.1186/1471-2164-12-72
Lu M, Liu S, Kumarsangaiah A (2017) Nucleosome positioning with fractal entropy increment of diversity in telemedicine. IEEE Access 6:33451–33459. https://doi.org/10.1109/ACCESS.2017.2779850
Magliery TJ, Regan L (2005) Sequence variation in ligand binding sites in proteins. BMC Bioinform 6:240. https://doi.org/10.1186/1471-2105-6-240
Mavrich TN, Ioshikhes IP, Venters BJ, Jiang C, Tomsho LP, Qi J, Schuster SC, Albert I, Pugh BF (2008) A barrier nucleosome model for statistical positioning of nucleosomes throughout the yeast genome. Genome Res. https://doi.org/10.1101/gr.078261.108
Meng Z, Shen H, Huang H (2018) Search result diversification on attributed networks via nonnegative matrix factorization. Inf Process Manag 54(6):1271–1291. https://doi.org/10.1016/j.ipm.2018.05.005
Peckham HE, Thurman RE, Fu Y, Stamatoyannopoulos JA, Noble WS, Struhl K, Weng Z (2007) Nucleosome positioning signals in genomic DNA. Genome Res. https://doi.org/10.1101/gr.6101007
Petralia F, Wang P, Yang J, Tu Z (2015) Integrative random forest for gene regulatory network inference. Bioinformatics 31(12):i197–i205. https://doi.org/10.1093/bioinformatics/btv268
Polishko A, Ponts N, Le Roch KG, Lonardi S (2012) Normal: accurate nucleosome positioning using a modified gaussian mixture model. Bioinformatics 28(12):i242–i249. https://doi.org/10.1093/bioinformatics/bts206
Rahman R, Otridge J, Pal R (2017) Integratedmrf: random forest-based framework for integrating prediction from different data types. Bioinformatics 33(9):1407–1410. https://doi.org/10.1093/bioinformatics/btw765
Sangaiah AK, Samuel OW, Li X (2017) Towards an efficient risk assessment in software projects—fuzzy reinforcement paradigm. Comput Electr Eng. https://doi.org/10.1016/j.compeleceng.2017.07.022
Sarosi G, Ugajin T (2016) Relative entropy of excited states in two dimensional conformal field theories. J High Energy Phys 2016:114. https://doi.org/10.1007/JHEP07(2016)114
Satchwell SC, Drew HR, Travers AA (1986) Sequence periodicities in chicken nucleosome core DNA. J Mol Biol 191(4):659–675. https://doi.org/10.1016/0022-2836(86)90452-3
Schones DE, Cui K, Cuddapah S, Roh TY, Barski A, Wang Z, Wei G, Zhao K (2008) Dynamic regulation of nucleosome positioning in the human genome. Cell 132(5):887–898. https://doi.org/10.1016/j.cell.2008.02.022
Segal E, Widom J (2009) Poly (DA: DT) tracts: major determinants of nucleosome organization. Curr Opin Struct Biol 19(1):65–71. https://doi.org/10.1016/j.sbi.2009.01.004
Shao LH, Li YM, Luo Y, Xi ZJ (2017) Quantum coherence quantifiers based on Renyi α-relative entropy. Commun Theor Phys 67(6):631–636. https://doi.org/10.1088/0253-6102/67/6/631
Sinoquet C (2018) A method combining a random forest-based technique with the modeling of linkage disequilibrium through latent variables, to run multilocus genome-wide association studies. BMC Bioinform 19:106. https://doi.org/10.1186/s12859-018-2054-0
Struhl K, Segal E (2013) Determinants of nucleosome positioning. Nat Struct Mol Biol 20:267–273. https://doi.org/10.1038/nsmb.2506
Taherzadeh G, Zhou Y, Liew AWC, Yang Y (2017) Structure-based prediction of protein-peptide binding regions using random forest. Bioinformatics 34(3):477–484. https://doi.org/10.1093/bioinformatics/btx614
Tahir M, Hayat M (2016) iNuc-STNC: a sequence-based predictor for identification of nucleosome positioning in genomes by extending the concept of saac and chou’s pseaac. Mol BioSyst 12(8):2587–2593. https://doi.org/10.1039/C6MB00221H
Tolstorukov MY, Choudhary V, Olson WK, Zhurkin VB, Park PJ (2008) nuScore: a web-interface for nucleosome positioning predictions. Bioinformatics 24(12):1456–1458. https://doi.org/10.1093/bioinformatics/btn212
Vacic V, Uversky VN, Dunker AK, Lonardi S (2007) Composition profiler: a tool for discovery and visualization of amino acid composition differences. BMC Bioinform 8:211. https://doi.org/10.1186/1471-2105-8-211
Vernikos GS, Parkhill J (2006) Interpolated variable order motifs for identification of horizontally acquired DNA: revisiting the salmonella pathogenicity islands. Bioinformatics 22(18):2196–2203. https://doi.org/10.1093/bioinformatics/btl369
Wan S, Mak MW, Kung SY (2013) GOASVM: a subcellular location predictor by incorporating term frequency gene ontology into the general form of Chou’s pseudo-amino acid composition. J Theor Biol 323:40–48. https://doi.org/10.1016/j.jtbi.2013.01.012
Wang K, Samudrala R (2006) Incorporating background frequency improves entropy-based residue conservation measures. BMC Bioinform 7:385. https://doi.org/10.1186/1471-2105-7-385
Woo S, Zhang X, Sauteraud R, Robert F, Gottardo R (2013) PING 2.0: an R/Bioconductor package for nucleosome positioning using next-generation sequencing data. Bioinformatics 29(16):2049–2050. https://doi.org/10.1093/bioinformatics/btt348
Xi L, Fondufe-Mittendorf Y, Xia L, Flatow J, Widom J, Wang JP (2010) Predicting nucleosome positioning using a duration Hidden Markov model. BMC Bioinform 11:346. https://doi.org/10.1186/1471-2105-11-346
Yasuda T, Sugasawa K, Shimizu Y, Iwai S, Shiomi T, Hanaoka F (2005) Nucleosomal structure of undamaged DNA regions suppresses the non-specific DNA binding of the XPC complex. DNA Repair 4(3):389–395. https://doi.org/10.1016/j.dnarep.2004.10.008
Yudong Z, Shuihua W, Ping S, Preetha P (2015) Pathological brain detection based on wavelet entropy and Hu moment invariants. Bio-Med Mater Eng 26(s1):S1283–S1290. https://doi.org/10.3233/BME-151426
Zhang YD, Wu LN (2008) Pattern recognition via PCNN and Tsallis entropy. Sensors 8(11):7518–7529. https://doi.org/10.3390/s8117518
Zhang Y, Wu L (2011) Optimal multi-level thresholding based on maximum Tsallis entropy via an artificial bee colony approach. Entropy 13(4):841–859. https://doi.org/10.3390/e13040841
Zhang Y, Gao X, Katayama S (2015) Weld appearance prediction with BP neural network improved by genetic algorithm during disk laser welding. J Manuf Syst 34:53–59. https://doi.org/10.1016/j.jmsy.2014.10.005
Zhang J, Hadj-Moussa H, Storey KB (2016) Current progress of high-throughput microRNA differential expression analysis and random forest gene selection for model and non-model systems: an R implementation. J Integr Bioinformatics 13(5):35–46. https://doi.org/10.1515/jib-2016-306
Zhang C, Li D, Sangaiah A (2017) Merger and acquisition target selection based on interval neutrosophic multigranulation rough sets over two universes. Symmetry 9(7):126. https://doi.org/10.3390/sym9070126
Zhang J, Peng W, Wang L (2018a) LeNup: learning nucleosome positioning from DNA sequences with improved convolutional neural networks. Bioinformatics 34(10):1705–1712. https://doi.org/10.1093/bioinformatics/bty003/4796955
Zhang C, Li D, Broumi S (2018b) Medical diagnosis based on single-valued neutrosophic probabilistic rough multisets over two universes. Symmetry 10(6):213. https://doi.org/10.3390/sym10060213
Acknowledgements
This research is funded by National Natural Science Foundation of China project with Grant No. 61502254, Program for Yong Talents of Science and Technology in Universities of Inner Mongolia Autonomous Region with Grant No. NJYT-18-B10, and Open Funds of Key Laboratory of Symbolic Computation and Knowledge Engineering of Ministry of Education with Grant No. 93K172018K07.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by A. K. Sangaiah, H. Pham, M.-Y. Chen, H. Lu, F. Mercaldo.
Rights and permissions
About this article
Cite this article
Lu, M., Liu, S. Nucleosome positioning based on generalized relative entropy. Soft Comput 23, 9175–9188 (2019). https://doi.org/10.1007/s00500-018-3602-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00500-018-3602-2