Abstract
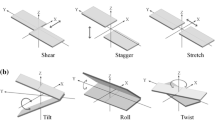
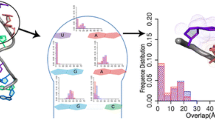
RNA contains different secondary structural motifs like pseudo-helices, hairpin loops, internal loops, etc. in addition to anti-parallel double helices and random coils. The secondary structures are mainly stabilized by base-pairing and stacking interactions between the planar aromatic bases. The hydrogen bonding strength and geometries of base pairs are characterized by six intra-base pair parameters. Similarly, stacking can be represented by six local doublet parameters. These dinucleotide step parameters can describe the quality of stacking between Watson–Crick base pairs very effectively. However, it is quite difficult to understand the stacking pattern for dinucleotides consisting of non canonical base pairs from these parameters. Stacking interaction is a manifestation of the interaction between two aromatic bases or base pairs and thus can be estimated best by the overlap area between the planar aromatic moieties. We have calculated base pair overlap between two consecutive base pairs as the buried van der Waals surface between them. In general, overlap values show normal distribution for the Watson–Crick base pairs in most double helices within a range from 45 to 50 Å2 irrespective of base sequence. The dinucleotide steps with non-canonical base pairs also are seen to have high overlap value, although their twist and few other parameters are rather unusual. We have analyzed hairpin loops of different length, bulges within double helical structures and pseudo-continuous helices using our algorithm. The overlap area analyses indicate good stacking between few looped out bases especially in GNRA tetraloop, which was difficult to quantitatively characterise from analysis of the base pair or dinucleotide step parameters. This parameter is also seen to be capable to distinguish pseudo-continuous helices from kinked helix junctions.





Similar content being viewed by others
Explore related subjects
Discover the latest articles and news from researchers in related subjects, suggested using machine learning.References
Neidle S (2002) Nucleic acid structure and recognition. Oxford University Press, Oxford
Leontis NB, Westhof E (2001) Geometric nomenclature and classification of RNA base pairs. RNA 7:499–512
Šponer JE, Špačková N, Leszczynski J, Šponer J (2005) Principles of RNA base pairing: structures and energies of the trans Watson − Crick/sugar edge base pairs. J Phys Chem B 109:11399–11410
Roy A, Panigrahi S, Bhattacharyya M, Bhattacharyya D (2008) Structure, stability, and dynamics of canonical and noncanonical base pairs: quantum chemical studies. J Phys Chem B 112:3786–3796
Halder S, Bhattacharyya D (2010) Structural stability of tandemly occurring noncanonical basepairs within double helical fragments: molecular dynamics studies of functional RNA. J Phys Chem B 114:14028–14040
Halder S, Bhattacharyya D (2012) Structural variations of single and tandem mismatches in RNA duplexes: a joint MD simulation and crystal structure database analysis. J Phys Chem B 116:11845–11856
Sarver M, Zirbel CL, Stombaugh J, Mokdad A, Leontis NB (2008) FR3D: finding local and composite recurrent structural motifs in RNA 3D structures. J Math Biol 56:215–252
Petrov AI, Zirbel CL, Leontis NB (2013) Automated classification of RNA 3D motifs and the RNA 3D Motif Atlas. RNA 19:1327–1340
Woese CR, Gutell RR (1989) Evidence for several higher order structural elements in ribosomal RNA. Proc Natl Acad Sci USA 86:3119–3122
Varani L, Hasegawa M, Spillantini MG, Smith MJ, Murrell JR, Ghetti B, Klug A, Goedert M, Varani G (1999) Structure of tau exon 10 splicing regulatory element RNA and destabilization by mutations of frontotemporal dementia and parkinsonism linked to chromosome 17. Proc Natl Acad Sci USA 96:8229–8234
Portmann S, Grimm S, Workman C, Usman N, Egli M (1996) Crystal structures of an A-form duplex with single-adenosine bulges and a conformational basis for site-specific RNA self-cleavage. Chem Biol 3:173–184
Hermann T, Patel DJ (1999) Stitching together RNA tertiary architectures. J Mol Biol 294:829–849
Hermann T, Westhof E (1999) Non-Watson–Crick base pairs in RNA–protein recognition. Chem Biol 6:R335–R343
Chastain M, Tinoco I (1991) Structural elements in RNA. Prog Nucleic Acid Res Mol Biol 41:131–177
Woese CR, Winkers S, Gutell RR (1990) Architecture of ribosomal RNA: constraints on the sequence of tetra-loops. Proc Natl Acad Sci USA 87:8467–8471
Heus HA, Pardi A (1991) Structural features that give rise to the unusual stability of RNA hairpins containing GNRA loops. Science 253:191–194
Jucker FM, Pardi A (1995) GNRA tetraloops make a U-turn. RNA 1:219–222
Leontis NB, Westhof E (2002) The annotation of RNA motifs. Comp Funct Genomics 3:518–524
Correll CC, Swinger K (2003) Common and distinctive features of GNRA tetraloops based on a GUAA tetraloop structure at 1.4Å resolution. RNA 9:355–363
Tuerk C, Gauss P, Thermes C, Groebe DR, Gayle M, Guild N, Stormo G, D’aubenton-Carafa Y, Uhlenbeck OC, Tinoco I Jr, Brody EN, Gold L (1988) CUUCGG hairpins: extraordinarily stable RNA secondary structures associated with various biochemical processes. Proc Natl Acad Sci USA 85:1364–1368
Cheong C, Varani G, Tinoco I Jr (1990) Solution structure of an unusually stable RNA hairpin, 5′GGAC(UUCG)GUCC. Nature 346:680–682
Baumruk V, Gouyette C, Huynh-Dinh T, Sun JS, Ghomi M (2001) Comparison between CUUG and UUCG Tetraloops: thermodynamic stability and structural features analyzed by UV absorption and vibrational spectroscopy. Nucleic Acids Res 29:4089–4096
Convery MA, Rowsell S, Stonehouse NJ, Ellington AD, Hirao I, Murray JB, Peabody DS, Phillips SE, Stockley PG (1998) Crystal structure of an RNA aptamer–protein complex at 2.8 Å resolution. Nat Struct Biol 5:133–139
Rowsell S, Stonehouse NJ, Convery MA, Adams CJ, Ellington AD, Hirao I, Peabody DS, Stockley PG, Phillips SEV (1998) Crystal structures of a series of RNA aptamers complexed to the same protein target. Nat Struct Biol 5:970–975
Klosterman PS, Hendrix DK, Tamura M, Holbrook SR, Brenner SE (2004) Three-dimensional motifs from the SCOR, structural classification of RNA database: extruded strands, base triples, tetraloops and U-turns. Nucleic Acids Res 32:2342–2352
Butcher SE, Dieckmann T, Feigon J (1997) Solution structure of the conserved 16S-like ribosomal RNA UGAA tetraloop. J Mol Biol 268:348–358
Lebars I, Lamontagne B, Yoshizawa S, Elela SA, Fourmy D (2001) Solution structure of conserved AGNN tetraloops: insights into Rnt1p RNA processing. EMBO J 20:7250–7258
Wu H, Yang PK, Butcher SE, Kang S, Chanfreau G, Feigon J (2001) A novel family of RNA tetraloop structures forms the recognition site for Saccharomyces cerevisiae RNaseIII. EMBO J 20:7240–7249
Woese CR, Gutell R, Gupta R, Noller HF (1983) Detailed analysis of the higher-order structure of 16S-like ribosomal ribonucleic acids. Microbiol Rev 4:621–669
Antao VP, Tinoco I Jr (1992) Thermodynamic parameters for loop formation in RNA and DNA hairpin tetraloops. Nucleic Acids Res 20:819–824
Proctor DJ, Schaak JE, Bevilacqua JM, Falzone CJ, Bevilacqua PC (2002) Isolation and characterization of a Family of Stable RNA Tetraloops with the motif YNMG That Participate in Tertiary Interactions. Biochemistry 41:12062–12075
Keating KS, Toor N, Pyle AM (2008) The GANC tetraloop: a novel motif in the group IIc intron structure. J Mol Biol 383:475–481
Zhao Q, Huang HC, Nagaswamy U, Xia Y, Gao X, Fox GE (2012) UNAC tetraloops: to what extent do they mimic GNRA tetraloops? Biopolymers 97:617–628
Melchers WJG, Zoll J, Tessari M, Bakhmutov DV, Gmyl AP, Agol VI, Heus HA (2006) A GCUA tetranucleotide loop found in poliovirus oriL by in vivo SELEX (un)expectedly forms a YNMG-like structure: extending the YNMG family with GYYA. RNA 12:1671–1682
Zwieb C (1992) Conformity of RNAs that interact with tetranucleotide loop binding proteins. Nucleic Acids Res 20:4397–4400
Sakamoto T, Morita S, Tabata K, Nakamura K, Kawai G (2002) Solution structure of a SRP 19 binding domain in human SRP RNA. J Biochem 132:177–182
Varani G, Cheong C, Tinoco I Jr (1991) Structure of an unusually stable RNA hairpin. Biochemistry 30:3280–3289
Lisi V, Major F (2007) A comparative analysis of the triloops in all high-resolution RNA structures reveals sequence–structure relationships. RNA 13:1537–1545
Lee JC, Cannone JJ, Gutell RR (2003) The lonepair triloop: a new motif in RNA structure. J Mol Biol 325:65–83
Halder S, Bhattacharyya D (2013) RNA structure and dynamics: a base pairing perspective. Prog Biophys Mol Biol 113:264–283
Richardson JS, Schneider B, Murray LW, Kapral GJ, Immormino RM, Headd JJ, Richardson DC, Ham D, Hershkovits E, Williams LD, Keating KS, Pyle AM, Micallef D, Westbrook J, Berman HM, RNA Ontology Consortium (2008) RNA backbone: consensus all-angle conformers and modular string nomenclature (an RNA Ontology Consortium contribution). RNA 14:465–481
Malathi R, Yathindra N (1985) Backbone conformation in nucleic acids: an analysis of local helicity through heminucleotide scheme and a proposal for a unified conformational plot. J Biomol Struct Dyn 3:127–144
Duarte CM, Pyle AM (1998) Stepping through an RNA structure: a novel approach to conformational analysis. J Mol Biol 284:1465–1478
Duarte CM, Wadley LM, Pyle AM (2003) RNA structure comparison, motif search and discovery using a reduced representation of RNA conformational space. Nucleic Acids Res 31:4755–4761
Wadley LM, Keating KS, Duarte CM, Pyle AM (2007) Evaluating and learning from RNA pseudotorsional space: quantitative validation of a reduced representation for RNA structure. J Mol Biol 372:942–957
Olson WK, Bansal M, Burley SK, Dickerson RE, Gerstein M, Harvey SC, Heinemann U, Lu XJ, Neidle S, Shakked Z, Sklenar H, Suzuki M, Tung CS, Westhof E, Wolfberger C, Berman HM (2001) A standard reference frame for the description of nucleic acid base-pair geometry. J Mol Biol 313:229–237
Samanta S, Mukherjee S, Chakrabarti J, Bhattacharyya D (2009) Structural properties of polymeric DNA from molecular dynamics simulations. J Chem Phys 130:115103
Marathe A, Bansal M (2011) An ensemble of B-DNA dinucleotide geometries lead to characteristic nucleosomal DNA structure and provide plasticity required for gene expression. BMC Struct Biol 11:1
Kailasam SK, Bhattacharyya D, Bansal M (2014) Sequence dependent variations in RNA duplex are related to non-canonical hydrogen bond interactions in dinucleotide steps. BMC Res Notes 7:83
Lu XJ, Olson WK (2003) 3DNA: a software package for the analysis, rebuilding and visualization of three-dimensional nucleic acid structures. Nucleic Acids Res 31:5108–5121
Gabb HA, Sanghani SR, Robert CH, Prévost C (1996) Finding and visualizing nucleic acid base stacking. J Mol Graph 14:6–11
Ray SS, Halder S, Kaypee S, Bhattacharyya D (2012) HD-RNAS: an automated hierarchical database of RNA structures. Front Genet 3:59
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Berman HM, Olson WK, Beveridge DL, Westbrook J, Gelbin A, Demeny T, Hsieh SH, Srinivasan AR, Schneider B (1992) The nucleic acid database: a comprehensive relational database of three-dimensional structures of nucleic acids. Biophys J 63:751–759
Das J, Mukherjee S, Mitra A, Bhattacharyya D (2006) Non-canonical base pairs and higher order structures in nucleic acids: crystal structure database analysis. J Biomol Struct Dyn 24:149–161
Bansal M, Bhattacharyya D, Ravi B (1995) NUPARM and NUCGEN: software for analysis and generation of sequence dependent nucleic acid structures. CABIOS 11:281–287
Mukherjee S, Bansal M, Bhattacharyya D (2006) Conformational specificity of non-canonical base pairs and higher order structures in nucleic acids: crystal structure database analysis. J Comput Aided Mol Des 20:629–645
Banerjee R, Sen M, Bhattacharyya D, Saha P (2003) The jigsaw puzzle model: search for conformational specificity in protein interiors. J Mol Biol 333:211–226
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM Jr, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
Chandrasekaran R, Wang M, He R-G, Puigjaner LC, Byler MA, Millane RP, Arnott S (1989) A re-examination of the crystal structure of A-DNA using fiber diffraction data. J Biomol Struct Dyn 6:1189–1202
Chandrasekaran R, Arnott S (1996) The structure of B-DNA in oriented fibers. J Biomol Struct Dyn 13:1015–1027
Arnott S, Hukins DWL, Dover SD (1972) Optimised parameters for RNA double-helices. Biochem Biophys Res Commun 48:1392–1399
Zhang F, Sahu B, Min H, MacDonald AH (2010) Bond structure of ABC-stacked graphene trilayers. Phys Rev B 82:035409
Mukherjee S, Kailasam SK, Bansal M, Bhattacharyya D (2014) Energy hyperspace for stacking interaction in AU/AU dinucleotide step: dispersion-corrected density functional theory study. Biopolymers 101:107–120
Calladine CR (1982) Mechanics of sequence-dependent stacking of bases in B-DNA. J Mol Biol 25:343–352
Acknowledgments
We are thankful to Prof. Manju Bansal, Indian Institute of Science, Bangalore, India for discussions. This work was partially supported by Department of Biotechnology, Govt. of India. PKP is thankful to National Institute of Pharmaceutical Education and Research, Kolkata for supports.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pingali, P.K., Halder, S., Mukherjee, D. et al. Analysis of stacking overlap in nucleic acid structures: algorithm and application. J Comput Aided Mol Des 28, 851–867 (2014). https://doi.org/10.1007/s10822-014-9767-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-014-9767-6