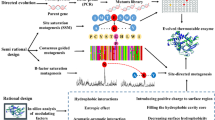
Abstract
Molecular level understanding of mutational effects on stability and activity of enzymes is challenging particularly when several point mutations are incorporated during the directed evolution experiments. In our earlier study, we have suggested the lack of consistency in the effect of point mutations incorporated during the initial generations of directed evolution experiments, towards conformational stabilization of B. subtilis lipase mutants of later generations. Here, we report that the cumulative point mutations incorporated in mutants 2M (with two point mutations) to 6M (with six point mutations) possibly do not retain their original stabilizing nature in the most thermostable 12M mutant (with 12 point mutations). We have carried out MD simulations using structures incorporating reversal of different sets of point mutations to assess their effect on the conformational stability and activity of 12M. Our analysis has revealed that reversal of certain point mutations in 12M had little effect on its conformational stability, suggesting that these mutations were probably inconsequential towards the thermostability of the 12M mutant. Interestingly these mutations involved evolutionarily conserved residues. On the other hand, some of the other point mutations incorporated in nonconserved regions, appeared to contribute significantly towards the conformational stability and/or activity of 12M. Based on the analysis of dynamics of in silico mutants generated using the consensus sequence, we identified experimentally verifiable residue positions to further increase the conformational stability and activity of the 12M mutant.













Similar content being viewed by others
References
Singh B, Bulusu G, Mitra A (2015) Understanding the thermostability and activity of Bacillus subtilis lipase mutants: insights from molecular dynamics simulations. J Phys Chem B 119:392–409. doi:10.1021/jp5079554
Polizzi KM, Bommarius AS, Broering JM, Chaparro-Riggers JF (2007) Stability of biocatalysts. Curr Opin Chem Biol 11:220–225. doi:10.1016/j.cbpa.2007.01.685
Yang H, Lu X, Liu L, Li J, Shin H, Chen RR, Du G, Chen J (2013) Fusion of an oligopeptide to the N terminus of an alkaline α-amylase from Alkalimonas amylolytica simultaneously improves the enzyme’s catalytic efficiency, thermal stability, and resistance to oxidation. Appl Environ Microbiol 79:3049–3058. doi:10.1128/AEM.03785-12
Blum JK, Ricketts MD, Bommarius AS (2012) Improved thermostability of AEH by combining B-FIT analysis and structure-guided consensus method. J Biotechnol 160:214–221. doi:10.1016/j.jbiotec.2012.02.014
Huang S-Y, Zhang Y-HP, Zhong J-J (2012) A thermostable recombinant transaldolase with high activity over a broad pH range. Appl Microbiol Biotechnol 93:2403–2410. doi:10.1007/s00253-011-3578-7
Rogers TA, Bommarius AS (2010) Utilizing Simple Biochemical Measurements to Predict Lifetime Output of Biocatalysts in Continuous Isothermal Processes. Chem Eng Sci 65:2118–2124. doi:10.1016/j.ces.2009.12.005
Xie Y, An J, Yang G, Wu G, Zhang Y, Cui L, Feng Y (2014) Enhanced enzyme kinetic stability by increasing rigidity within the active site. J Biol Chem 289:7994–8006. doi:10.1074/jbc.M113.536045
Sanchez-Ruiz JM (2010) Protein kinetic stability. Biophys Chem 148:1–15. doi:10.1016/j.bpc.2010.02.004
Vemparala S, Mehrotra S, Balaram H (1814) Role of loop dynamics in thermal stability of mesophilic and thermophilic adenylosuccinate synthetase: a molecular dynamics and normal mode analysis study. Biochim Biophys Acta 2011:630–637. doi:10.1016/j.bbapap.2011.03.012
Tavernelli I, Cotesta S, Di Iorio EE (2003) Protein dynamics, thermal stability, and free-energy landscapes: a molecular dynamics investigation. Biophys J 85:2641–2649. doi:10.1016/S0006-3495(03)74687-6
Tiberti M, Papaleo E (2011) Dynamic properties of extremophilic subtilisin-like serine-proteases. J Struct Biol 174:69–83. doi:10.1016/j.jsb.2011.01.006
Green SM, Shortle D (1993) Patterns of nonadditivity between pairs of stability mutations in staphylococcal nuclease. Biochemistry 32:10131–10139
LiCata VJ, Ackers GK (1995) Long-range, small magnitude nonadditivity of mutational effects in proteins. Biochemistry 34:3133–3139
Matsuura T, Yomo T, Trakulnaleamsai S, Ohashi Y, Yamamoto K, Urabe I (1998) Nonadditivity of mutational effects on the properties of catalase I and its application to efficient directed evolution. Protein Eng 11:789–795. doi:10.1093/protein/11.9.789
Istomin AY, Gromiha MM, Vorov OK, Jacobs DJ, Livesay DR (2008) New insight into long-range nonadditivity within protein double-mutant cycles. Proteins. 70:915–924. doi:10.1002/prot.21620
Reetz MT (2013) The Importance of Additive and Non-Additive Mutational Effects in Protein Engineering. Angew Chem Int Ed 52:2658–2666. doi:10.1002/anie.201207842
Bank C, Hietpas RT, Jensen JD, Bolon DNA (2015) A systematic survey of an intragenic epistatic landscape. Mol Biol Evol 32:229–238. doi:10.1093/molbev/msu301
Ahmad S, Rao NM (2009) Thermally denatured state determines refolding in lipase: Mutational analysis. Protein Sci 18:1183–1196. doi:10.1002/pro.126
Ahmad S, Kumar V, Ramanand KB, Rao NM (2012) Probing protein stability and proteolytic resistance by loop scanning: a comprehensive mutational analysis. Protein Sci 21:433–446. doi:10.1002/pro.2029
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Kamal MZ, Ahmad S, Molugu TR, Vijayalakshmi A, Deshmukh MV, Sankaranarayanan R, Rao NM (2011) In vitro evolved non-aggregating and thermostable lipase: structural and thermodynamic investigation. J Mol Biol 413:726–741. doi:10.1016/j.jmb.2011.09.002
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. doi:10.1002/jcc.20084
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, van der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854. doi:10.1093/bioinformatics/btt055
Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, Shaw DE (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins. 78:1950–1958. doi:10.1002/prot.22711
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79:926–935. doi:10.1063/1.445869
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J. Chem. Phys. 126:14101. doi:10.1063/1.2408420
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190. doi:10.1063/1.328693
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J. Chem. Phys. 103:8577–8593. doi:10.1063/1.470117
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472. doi:10.1002/(SICI)1096-987X(199709)18:12<1463:AID-JCC4>3.0.CO;2-H
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637. doi:10.1002/bip.360221211
Warren L. DeLano “The PyMOL Molecular Graphics System.” DeLano Scientific LLC, San Carlos, CA, USA. http://www.pymol.org (n.d.)
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(33–38):27–28
Ichiye T, Karplus M (1991) Collective motions in proteins: a covariance analysis of atomic fluctuations in molecular dynamics and normal mode simulations. Proteins. 11:205–217. doi:10.1002/prot.340110305
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2015) CDD: NCBI’s conserved domain database. Nucleic Acids Res 43:D222–D226. doi:10.1093/nar/gku1221
Boratyn GM, Schäffer AA, Agarwala R, Altschul SF, Lipman DJ, Madden TL (2012) Domain enhanced lookup time accelerated BLAST. Biol. Direct. 7:12. doi:10.1186/1745-6150-7-12
Srivastava A, Sinha S (2014) Thermostability of In Vitro Evolved Bacillus subtilis Lipase A: a Network and Dynamics Perspective. PLoS ONE 9:e102856. doi:10.1371/journal.pone.0102856
Senthilkumar B, Meshachpaul D, Sethumadhavan R, Rajasekaran R (2015) Selection of effective and highly thermostable Bacillus subtilis lipase A template as an industrial biocatalyst-A modern computational approach. Front. Biol. 10:508–519. doi:10.1007/s11515-015-1379-6
Tiberti M, Invernizzi G, Lambrughi M, Inbar Y, Schreiber G, Papaleo E (2014) PyInteraph: a framework for the analysis of interaction networks in structural ensembles of proteins. J Chem Inf Model 54:1537–1551. doi:10.1021/ci400639r
Ahmad S, Kamal MZ, Sankaranarayanan R, Rao NM (2008) Thermostable Bacillus subtilis lipases: in vitro evolution and structural insight. J Mol Biol 381:324–340. doi:10.1016/j.jmb.2008.05.063
Acharya P, Rajakumara E, Sankaranarayanan R, Rao NM (2004) Structural basis of selection and thermostability of laboratory evolved Bacillus subtilis lipase. J Mol Biol 341:1271–1281. doi:10.1016/j.jmb.2004.06.059
Prajapati RS, Das M, Sreeramulu S, Sirajuddin M, Srinivasan S, Krishnamurthy V, Ranjani R, Ramakrishnan C, Varadarajan R (2007) Thermodynamic effects of proline introduction on protein stability. Proteins. 66:480–491. doi:10.1002/prot.21215
Ramachandran GN, Ramakrishnan C, Sasisekharan V (1963) Stereochemistry of polypeptide chain configurations. J Mol Biol 7:95–99. doi:10.1016/S0022-2836(63)80023-6
Lehmann M, Pasamontes L, Lassen SF, Wyss M (2000) The consensus concept for thermostability engineering of proteins. Biochim Biophys Acta 1543:408–415
Lehmann M, Wyss M (2001) Engineering proteins for thermostability: the use of sequence alignments versus rational design and directed evolution. Curr Opin Biotechnol 12:371–375
Lehmann M, Loch C, Middendorf A, Studer D, Lassen SF, Pasamontes L, van Loon APGM, Wyss M (2002) The consensus concept for thermostability engineering of proteins: further proof of concept. Protein Eng 15:403–411. doi:10.1093/protein/15.5.403
Godoy-Ruiz R, Ariza F, Rodriguez-Larrea D, Perez-Jimenez R, Ibarra-Molero B, Sanchez-Ruiz JM (2006) Natural selection for kinetic stability is a likely origin of correlations between mutational effects on protein energetics and frequencies of amino acid occurrences in sequence alignments. J Mol Biol 362:966–978. doi:10.1016/j.jmb.2006.07.065
Pey AL, Rodriguez-Larrea D, Bomke S, Dammers S, Godoy-Ruiz R, Garcia-Mira MM, Sanchez-Ruiz JM (2008) Engineering proteins with tunable thermodynamic and kinetic stabilities. Proteins. 71:165–174. doi:10.1002/prot.21670
Bloom JD, Glassman MJ (2009) Inferring stabilizing mutations from protein phylogenies: application to influenza hemagglutinin. PLoS Comput Biol 5:e1000349. doi:10.1371/journal.pcbi.1000349
Jäckel C, Bloom JD, Kast P, Arnold FH, Hilvert D (2010) Consensus protein design without phylogenetic bias. J Mol Biol 399:541–546. doi:10.1016/j.jmb.2010.04.039
Cole MF, Gaucher EA (2011) Exploiting models of molecular evolution to efficiently direct protein engineering. J Mol Evol 72:193–203. doi:10.1007/s00239-010-9415-2
Acknowledgments
B.S. and A.M. thank DBT, Government of India Project BT/PR-14715/PBD/16/903/2010 for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10822_2016_9978_MOESM1_ESM.doc
Supplementary material 1: MD simulation results of all the systems, supporting texts, figures and tables discussed in the manuscript are available online as supplementary material. (DOC 953 kb)
Rights and permissions
About this article
Cite this article
Singh, B., Bulusu, G. & Mitra, A. Effects of point mutations on the thermostability of B. subtilis lipase: investigating nonadditivity. J Comput Aided Mol Des 30, 899–916 (2016). https://doi.org/10.1007/s10822-016-9978-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-9978-0