Abstract
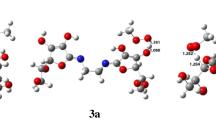
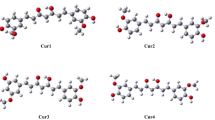
During a respiratory burst the enzyme myeloperoxidase generates significant amounts of hypohalous acids (HOX, X = Cl and Br) in order to inflict oxidative damage upon invading pathogens. However, excessive production of these potent oxidants is associated with numerous inflammatory diseases. It has been suggested that the endogenous antioxidant carnosine is an effective HOCl scavenger. Recent computational and experimental studies suggested that an intramolecular Cl+ transfer from the imidazole ring to the terminal amine might play an important role in the antioxidant activity of carnosine. Based on high-level ab initio calculations, we propose a similar reaction mechanism for the intramolecular Br+ transfer in carnosine. These results suggest that carnosine may be an effective HOBr scavenger. On the basis of the proposed reaction mechanism, we proceed to design systems that share similar structural features to carnosine but with enhanced HOX scavenging capabilities for X = Cl and Br. We find that (i) elongating the β-alanyl-glycyl side chain by one carbon reduces the reaction barriers by up to 44%, and (ii) substituting the imidazole ring with strong electron-donating groups reduces the reaction barriers by similar amounts. We also show that the above structural and electronic effects are largely additive. In an antioxidant candidate that involves both of these effects the reaction barriers are reduced by 71%.







Similar content being viewed by others
References
van Dalen C, Whitehouse M, Winterbourn C, Kettle A (1997) Biochem J 327:487
Slungaard A, Mahoney JR (1991) J Biol Chem 266:4903
Thomas EL, Fishman M (1986) J Biol Chem 261:9694
van der Veen BS, de Winther MP, Heeringa P (2009) Antioxid Redox Signal 11:2899
Klebanoff SJ (2005) J Leukoc Biol 77:598
Valko M, Leibfritz D, Moncol J, Cronin MT, Mazur M, Telser J (2007) Int J Biochem Cell Biol 39:44
Yap YW, Whiteman M, Cheung NS (2007) Cell Signal 19:219
Malle E, Marsche G, Arnhold J, Davies MJ (2006) Biochim Biophys Acta 1761:392
Balaban RS, Nemoto S, Finkel T (2005) Cell 120:483
Stocker R, Keaney JF Jr (2004) Physiol Rev 84:1381
Wu W, Samoszuk MK, Comhair SAA, Thomassen MJ, Farver CF, Dweik RA, Kavuru MS, Erzurum SC, Hazen SL (2000) J Clin Invest 105:1455
Aldridge RE, Chan T, van Dalen CJ, Senthilmohan R, Winn M, Venge P, Town GI, Kettle AJ (2002) Free Rad Biol Med 33:847
Spry CJF (1988) Eosinophils: a comprehensive review, and guide to the scientific and medical literature. Oxford University Press, Oxford
Kaliyeva L, Zhumagali S, Akhmetova N, Karton A, O’Reilly RJ (2017) Int J Quantum Chem 117:e25319
O’Reilly RJ, Karton A (2016) Int J Quantum Chem 116:52
O’Reilly RJ, Karton A, Radom. L (2013) J Phys Chem A 117:460
O’Reilly RJ, Karton A, Radom L (2012) Int J Quantum Chem 112:1862
O’Reilly RJ, Karton A, Radom L (2011) J Phys Chem A 115:5496
Sivey JD, Howell SC, Bean DJ, McCurry DL, Mitch WA, Wilson CJ (2013) Biochemistry 52:1260
Pattison DI, Hawkins CL, Davies MJ (2009) Chem Res Toxicol 22:807
Hawkins CL (2009) Free Radic Res 43:1147
Davies MJ, Hawkins CL, Pattison DI, Rees MD (2008) Antioxid Redox Signal 10:1199
Pattison DI, Davies MJ (2006) Curr Med Chem 13:3271
Pattison DI, Davies MJ (2005) Biochemistry 44:7378
Hawkins CL, Pattison DI, Davies MJ (2003) Amino Acids 25:259
Pattison DI, Davies MJ (2004) Biochemistry 43:4799
Thomas EL, Bozeman PM, Jefferson MM, King CC (1995) J Biol Chem 270:2906
Carr AC, Winterbourn CC, van den Berg JJ (1996) Arch Biochem Biophys 327:227
Vissers M, Carr A, Chapman A (1998) Biochem J 330:131
Henderson JP, Byun J, Mueller DM, Heinecke JW (2001) Biochemistry 40:2052
Henderson JP, Byun J, Williams MV, McCormick ML, Parks WC, Ridnour LA, Heinecke JW (2001) Proc Natl Acad Sci USA 98:1631
Drozak J, Veiga-da-Cunha M, Vertommen D, Stroobant V, Van Schaftingen E (2010) J Biol Chem 285:9346
Quinn PJ, Boldyrev AA, Formazuyk VE (1992) Mol Aspects Med 13:379
Hipkiss AR (2009) Adv Food Nutr Res 57:87
Hipkiss AR, Worthington VC, Himsworth DTJ, Herwig W (1998) Biochim Biophys Acta 1380:46
Pattison DI, Davies MJ (2006) Biochemistry 45:8152
Karton A, O’Reilly RJ, Pattison DI, Davies MJ, Radom L (2012) J Am Chem Soc 134:19240
Lee C, Yang W, Parr RG (1988) Phys Rev B 37:785
Becke AD (1993) J Chem Phys 98:5648
Stephens PJ, Devlin FJ, Chabalowski CF, Frisch MJ (1994) J Phys Chem 98:11623
Grimme S, Ehrlich S, Goerigk L (2011) J Comput Chem 32:1456
Grimme S, Antony J, Ehrlich S, Krieg H (2010) J Chem Phys 132:154104
Grimme S (2011) WIREs Comput Mol Sci 1:211
Becke AD, Johnson ER (2005) J Chem Phys 123:154101
Marenich AV, Cramer CJ, Truhlar DG (2009) J Phys Chem B 113:6378
Gonzalez C, Schlegel HB (1989) J Chem Phys 90:2154
Gonzalez C, Schlegel HB (1990) J Phys Chem 94:5523
Hanwell MD, Curtis DE, Lonie DC, Vandermeersch T, Zurek E, Hutchison GR (2012) J Cheminform 4:17
Curtiss LA, Redfern PC, Raghavachari K (2007) J Chem Phys 127:124105
Curtiss LA, Redfern PC, Raghavachari K (2011) Wiley Interdiscip Rev Comput Mol Sci 1:810
Karton A (2016) Wiley Interdiscip Rev Comput Mol Sci 6:292
Curtiss LA, Redfern PC, Raghavachari K (2005) J Chem Phys 123:124107
Curtiss LA, Redfern PC, Raghavachari K (2010) Chem Phys Lett 499:168
Karton A, O’Reilly RJ, Radom L (2012) J Phys Chem A 116:4211
Karton A, Goerigk L (2015) J Comput Chem 36:622
Yu L-J, Sarrami F, O’Reilly RJ, Karton A (2015) Chem Phys 458:1
Cossi M, Rega N, Scalmani G, Barone V (2003) J Comput Chem 24:669
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA et al (2009) Gaussian 09, Revision E.01, Gaussian Inc., Wallingford, CT
Cioslowski J (1989) J Am Chem Soc 111:8333
De Proft F, Martin JML., Geerlings P (1996) Chem Phys Lett 250:393
Diez RP, Baran EJ (2003) J Mol Struct 621:245
Hansch C, Leo A, Taft RW (1991) Chem Rev 91:165
Acknowledgements
This work is dedicated to our colleague and friend Dr. Ming Wen Shi, who tragically passed away earlier this year. This research was undertaken with the assistance of resources from the National Computational Infrastructure (NCI), which is supported by the Australian Government. We also acknowledge the system administration support provided by the Faculty of Science at the University of Western Australia to the Linux cluster of the Karton group. We gratefully acknowledge the provision of an Australian Postgraduate Award (to F.S.), and an Australian Research Council (ARC) Discovery Early Career Researcher Award (to A.K., Project No. DE140100311). We would also like to thank the reviewers of the manuscript for their valuable comments and suggestions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sarrami, F., Yu, LJ. & Karton, A. Computational design of bio-inspired carnosine-based HOBr antioxidants. J Comput Aided Mol Des 31, 905–913 (2017). https://doi.org/10.1007/s10822-017-0060-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-017-0060-3