Abstract
Purpose
In the current standard of care, real-time transrectal ultrasound (TRUS) is commonly used for prostate brachytherapy guidance. As TRUS provides limited soft tissue contrast, segmenting the prostate gland in TRUS images is often challenging and subject to inter-observer and intra-observer variability, especially at the base and apex where the gland boundary is hard to define. Magnetic resonance imaging (MRI) has higher soft tissue contrast allowing the prostate to be contoured easily. In this paper, we aim to show that prostate segmentation in TRUS images informed by MRI priors can improve on prostate segmentation that relies only on TRUS images.
Methods
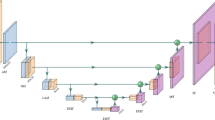
First, we compare the TRUS-based prostate segmentation used in the treatment of 598 patients with a high-quality MRI prostate atlas and observe inconsistencies at the apex and base. Second, motivated by this finding, we propose an alternative TRUS segmentation technique that is fully automatic and uses MRI priors. The algorithm uses a convolutional neural network to segment the prostate in TRUS images at mid-gland, where the gland boundary can be clearly seen. It then reconstructs the gland boundary at the apex and base with the aid of a statistical shape model built from an MRI atlas of 78 patients.
Results
Compared to the clinical TRUS segmentation, our method achieves similar mid-gland segmentation results in the 598-patient database. For the seven patients who had both TRUS and MRI, our method achieved more accurate segmentation of the base and apex with the MRI segmentation used as ground truth.
Conclusion
Our results suggest that utilizing MRI priors in TRUS prostate segmentation could potentially improve the performance at base and apex.




Similar content being viewed by others
References
Morris WJ, Keyes M, Palma D, Spadinger I, McKenzie MR, Agranovich A, Pickles T, Liu M, Kwan W, Wu J, Berthelet E, Pai H (2009) Population-based study of biochemical and survival outcomes after permanent 125I brachytherapy for low- and intermediate-risk prostate cancer. Urology 73(4):860–865
Badiei S, Salcudean SE, Varah J, Morris WJ (2006) Prostate segmentation in 2D ultrasound images using image warping and ellipse fitting. In: International conference on medical image computing and computer-assisted intervention. Springer, Berlin, pp 17–24
Soumya G, Arnau O, Robert M, Xavier L, Joan CV, Jordi F, Jhimli M, Dsir S, Fabrice M (2012) A survey of prostate segmentation methodologies in ultrasound, magnetic resonance and computed tomography images. Comput Methods Programs Biomed 108(1):262–287
Pathak SD, Haynor DR, Kim Y (2000) Edge-guided boundary delineation in prostate ultrasound images. IEEE Trans Med Imaging 19(12):1211–1219
Gong L, Pathak SD, Haynor DR, Cho PS, Kim Y (2004) Parametric shape modeling using deformable superellipses for prostate segmentation. IEEE Trans Med Imaging 23(3):340–349
Shen D, Zhan Y, Davatzikos C (2003) Segmentation of prostate boundaries from ultrasound images using statistical shape model. IEEE Trans Med Imaging 22(4):539–551
Mahdavi SS, Chng N, Spadinger I, Morris WJ, Salcudean SE (2011) Semi-automatic segmentation for prostate interventions. Med Image Anal 15(2):226–237
Mahdavi SS, Spadinger I, Chng N, Salcudean SE, Morris WJ (2013) Semiautomatic segmentation for prostate brachytherapy: dosimetric evaluation. Brachytherapy 12(1):65–76
Abolmaesumi P, Sirouspour MR (2004) An interacting multiple model probabilistic data association filter for cavity boundary extraction from ultrasound images. IEEE Trans Med Imaging 23(6):772–784
Nouranian S, Mahdavi SS, Spadinger I, Morris WJ, Salcudean SE, Abolmaesumi P (2015) A multi-atlas based segmentation framework for prostate brachytherapy. IEEE Trans Med Imaging 34(4):950–961
Nouranian S, Ramezani M, Spadinger I, Morris WJ, Salcudean SE, Abolmaesumi P (2016) Learning-based multi-label segmentation of transrectal ultrasound images for prostate brachytherapy. IEEE Trans Med Imaging 35(3):921–931
Anas EM, Nouranian S, Mahdavi SS, Spadinger I, Morris WJ, Salcudean SE, Mousavi P, Abolmaesumi P (2017) Clinical target-Volume delineation in prostate brachytherapy using residual neural networks. In: International conference on medical image computing and computer-assisted intervention. Springer, Berlin, pp 365–373
Martin S, Daanen V, Troccaz S (2008) Atlas-based prostate segmentation using an hybrid registration. Int J Comput Assist Radiol Surg 3(6):485–492
Rahmouni A, Yang A, Tempany CM, Frenkel T, Epstein J, Walsh P, Leichner PK, Ricci C, Zerhouni E (1992) Accuracy of in-vivo assessment of prostatic volume by MRI and transrectal ultrasonography. J Comput Assist Tomogr 16(6):935–940
Lee JS, Chung BH (2007) Transrectal ultrasound versus magnetic resonance imaging in the estimation of prostate volume as compared with radical prostatectomy specimens. Urol Int 78(4):323–327
Reynier C, Troccaz J, Fourneret P, Dusserre A, Gay-Jeune C, Descotes JL, Bolla M, Giraud JY (2004) MRI/TRUS data fusion for prostate brachytherapy. Preliminary results. Med Phys 31(6):1568–1575
Khallaghi S, Snchez CA, Rasoulian A, Nouranian S, Romagnoli C, Abdi H, Chang SD, Black PC, Goldenberg L, Morris WJ, Spadinger I, Fenster A, Ward A, Fels S, Abolmaesumi P (2015) Statistical biomechanical surface registration: application to MR–TRUS fusion for prostate interventions. IEEE Trans Med Imaging 34(12):2535–2549
Litjens G, Toth R, van de Ven W, Hoeks C, Kerkstra S, van Ginneken B, Vincent G, Guillard G, Birbeck N, Zhang J, Strand R, Malmberg F, Ou Y, Davatzikos C, Kirschner M, Jung F, Yuan J, Qiu W, Gao Q, Edwards PE, Maan B, van der Heijden F, Ghose S, Mitra J, Dowling J, Barratt D, Huisman H, Madabhushi A (2014) Evaluation of prostate segmentation algorithms for MRI: the PROMISE12 challenge. Med Image Anal 18(2):359–373
Ronneberger O, Fischer P, Brox T (2015) U-net: convolutional networks for biomedical image segmentation. In: International conference on medical image computing and computer-assisted intervention. Springer, Berlin, pp 234–241
Milletari F, Navab N, Ahmadi S (2016) V-net: fully convolutional neural networks for volumetric medical image segmentation. In: IEEE international conference on 3D vision, pp 565–571
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. In: Proceedings of the 3rd international conference on learning representations. arXiv:1409.1556
Xu B, Wang N, Chen T, Li M (2015) Empirical evaluation of rectified activations in convolutional network. arXiv:1505.00853
Gao H, Zhuang L, Kilian QW, Laurens van der M (2017) Densely connected convolutional networks. In: IEEE conference on computer vision and pattern recognition, pp 2261–2269
Havaei M, Axel Davy A, David Warde-Farley D, Antoine Biard A, Aaron Courville A, Yoshua Bengio Y, Chris Pal C, Pierre-Marc Jodoin P, Hugo Larochelle H (2017) Brain tumor segmentation with deep neural networks. Med Image Anal 35:18–31
Salehi S, Erdogmus D, Gholipour A (2017) Tversky loss function for image segmentation using 3D fully convolutional deep networks. In: International workshop on machine learning in medical, imaging, pp 379–387
Kingma DP, Ba J (2015) Adam: a method for stochastic optimization. In: Proceedings of the 3rd international conference on learning representations. arXiv:1412.6980
Rasoulian A, Rohling R, Abolmaesumi P (2012) Group-wise registration of point sets for statistical shape models. IEEE Trans Med Imaging 31(11):2025–2034
Blanz V, Mehl A, Vetter T, Seidel HP (2004) A statistical method for robust 3D surface reconstruction from sparse data. In: Proceedings of 2nd international symposium on 3D data processing, visualization and transmission, pp 293–300
Myronenko A, Song X (2010) Point set registration: coherent point drift. IEEE Trans Pattern Anal Mach Intell 32(12):2262–2275
Acknowledgements
This work was funded by Natural Sciences and Engineering Research Council of Canada (NSERC), the Canadian Institutes of Health Research (CIHR), and the Prostate Cancer Canada (PCC). We would like to thank the support from the Charles Laszlo Chair in Biomedical Engineering held by Professor Salcudean. The authors also gratefully acknowledge the help from physicians and staff at the Vancouver Cancer Centre who have contributed to this project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Zeng, Q., Samei, G., Karimi, D. et al. Prostate segmentation in transrectal ultrasound using magnetic resonance imaging priors. Int J CARS 13, 749–757 (2018). https://doi.org/10.1007/s11548-018-1742-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-018-1742-6