Abstract
Purpose
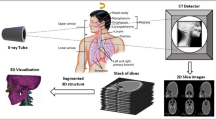
In computer-assisted diagnosis for orthopedic treatment, 3D reconstruction of bones is critical. Traditional 3D imaging technologies like CT and MRI have been proposed, but their high radiation dose and the requirements for lying postures could impact the accuracy of reconstructed bones and diagnosis results. Meanwhile, methods based on bone contours always depend on prior knowledge and lack precise bone segmentation methods. To address these issues, a bone reconstruction method based on multi-views of contours is proposed, as well as a hybrid CNN-Transformer approach for bone contours segmentation.
Methods
A four-step strategy is introduced including segmenting bone contours from X-ray images, calculating 3D sparse, dense point clouds based on contours, and reconstructing surface. The Trans-DetSeg approach for interest regions detection and bone segmentation is proposed for accurate contours. Besides, the mathematical description of mapping relationships between contours in different views of X-ray images is provided. Then, bone sparse and dense point clouds are generated subsequently. Based on dense point clouds and the power crust method, realistic bone models are reconstructed.
Results
Evaluated on 301 bone X-ray images and by considering p-value < 0.05, the proposed Trans-Detseg approach performed better with Dice Similarity Coefficient of 0.949 and Hausdorff Distance of 26.17 than three state-of-the-art models. Furthermore, the accuracy of the bone 3D reconstruction was investigated in three tibia cases and the proposed method was verified based on comparisons of results and CT data. It was proved that increased views of X-ray images could reduce the Average Surface Distance and perfect the structure information of reconstructed bones.
Conclusion
A new method for bone 3D reconstruction based on segmented bone contours on multi-views of X-ray images has been developed. Besides, a hybrid CNN-Transformer approach is introduced to segment bone contours. Evaluations proved the efficiency and accuracy of the proposed bone 3D reconstruction method.













Similar content being viewed by others
References
Hosseinian S, Arefi H (2015) 3D Reconstruction from multi-views medical x-ray images - review and evaluation of existing methods. In: The international archives of the photogrammetry, remote sensing and spatial information sciences, 40
Wang J, Ye M, Liu Z, Wang C (2009) Precision of cortical bone reconstruction based on 3D CT scans. Comput Med Imaging Graph 33(3):235–241
Radzi S, Cowin G, Robinson M, Pratap J, Volp A, Schuetz MA, Schmutz B (2014) Metal artifacts from titanium and steel screws in CT, 1.5 T and 3T MR images of the tibial Pilon: a quantitative assessment in 3D. Quant Imag Med Surg 4(3):163
Baka N, Kaptein BL, de Bruijne M, van Walsum T, Giphart JE, Niessen WJ, Lelieveldt BPF (2011) 2D–3D shape reconstruction of the distal femur from stereo X-ray imaging using statistical shape models. Med Image Anal 15(6):840–850
Murat G, Shimada K (2004) Three-dimensional bone shape reconstruction from X-ray images using hierarchical free-form deformation and nonlinear optimization. Int Congr Ser 1268:1291–1291
Fernandez JW, Mithraratne P, Thrupp SF, Tawhai MH, Hunter PJ (2004) Anatomically based geometric modelling of the musculo-skeletal system and other organs. Biomech Model Mechanobiol 2(3):139–155
Schmutz B, Reynolds KJ, Slavotinek JP (2009) Customization of a generic 3D model of the distal femur using diagnostic radiographs. J Med Eng Technol 32(2):156–161
Messmer P, Long G, Suhm N, Regazzoni P, Jacob AL (2001) Volumetric model determination of the tibia based on 2D radiographs using a 2D/3D database. Comput Aided Surg 6(4):183–194
Kohta I, Hosoda K, Shimizu M, Ikemoto S, Kume S, Nagura T, Imanishi N, Aiso S, Jinzaki M, Ogihara N (2015) Direct assessment of 3D foot bone kinematics using biplanar X-ray fluoroscopy and an automatic model registration method. J Foot Ankle Res 8(1):1–10
Caponetti L, Fanelli A (1993) Computer-aided simulation for bone surgery. IEEE Comput Graphics Appl 13(6):86–92
Caponetti L, Fanelli A (1990) 3D bone reconstruction from two X-ray views. In: Proceedings of the twelfth annual international conference of the IEEE engineering in medicine and biology society. pp 208–210
Nikkhade-Dehkordi B, Bro-Nielsen M, Darvann T, Gramkow C, Egund N, Hermann K (1996) 3D reconstruction of the femoral bone using two X-ray images from orthogonal views. Comput-Assist Radiol Int Symp Comput Commun Syst Image Guided Diag Therapy 1124:1066–1068
Mitulescu A, Skalli W, Mitton D, De Guise J (2002) Three-dimensional surface rendering reconstruction of scoliotic vertebrae using a non-stereo-corresponding points technique. Eur Spine J 11(4):344–352
Garg B, Mehta N, Bansal T, Malhotra R (2020) EOS® imaging: Concept and current applications in spinal disorders. J Clinical Orthopaed Trauma 11(5):786–793
Senthilkumaran N, Vaithegi S (2016) Image segmentation by using thresholding techniques for medical images. Comput Sci Eng Int J 6(1):1–13
Naz S, Majeed H, Irshad H (2010) Image segmentation using fuzzy clustering: A survey. In: 2010 6th international conference on emerging technologies. pp 181–186
Preetha MMSJ, Suresh LP, Bosco MJ (2012) Image segmentation using seeded region growing. In: 2012 international conference on computing, electronics and electrical technologies pp 576–583
Wang R, Lei T, Cui R, Zhang B, Meng H, Nandi AK (2020) Medical image segmentation using deep learning: A survey. IET Image Proc 16(5):1243–1267
Wang S, Zhou M, Liu Z, Liu Z, Gu D, Zang Y, Dong D, Gevaert O, Tian J (2017) Central focused convolutional neural networks: Developing a data-driven model for lung nodule segmentation. Med Image Anal 40:172–183
Jonathan L, Shelhamer E, Darrell T (2015) Fully convolutional networks for semantic segmentation. In: Proceedings of the IEEE conference on computer vision and pattern recognition pp 3431–3440
Ronneberger O, Fischer P, Brox T (2015) U-net: Convolutional networks for biomedical image segmentation. In: International conference on medical image computing and computer-assisted intervention pp 234–241
Lian S, Luo Z, Zhong Z, Lin X, Su S, Li S (2018) Attention guided U-Net for accurate iris segmentation. J Vis Commun Image Represent 56:296–304
Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, Lin S, Guo B (2021) Swin transformer: Hierarchical vision transformer using shifted windows. In: Proceedings of the IEEE/CVF international conference on computer vision pp 10012–10022
Valanarasu JMJ, Oza P, Hacihaliloglu I, Patel VM (2021) Medical transformer: Gated axial-attention for medical image segmentation. In: International conference on medical image computing and computer-assisted intervention pp 36–46
Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez AN, Kaiser L, Polosukhin I (2017) Attention is all you need. Adv Neural Inf Process Syst 5998–6008
Parmar N, Vaswani A, Uszkoreit J, Kaiser L, Shazeer N, Ku A, Tran D (2018) Image transformer. In: International conference on machine learning pp 4055–4064
Radford A, Kim JW, Hallacy C, Ramesh A, Goh G, Agarwal S, Sastry G, Askell A, Mishkin P, Clark J, Krueger G (2021) Learning transferable visual models from natural language supervision. In: International conference on machine learning pp 8748–8763
Amiri M, Brooks R, Behboodi B, Rivaz H (2020) Two-stage ultrasound image segmentation using U-Net and test time augmentation. Int J Comput-Assisted Radiol Surg 15(6):981–988
Carion N, Massa F, Synnaeve G, Usunier N, Kirillov A, Zagoruyko S (2020) End-to-end object detection with transformers. In: European conference on computer vision pp 213–229
Gamage P, Xie SQ, Patrice D, Xu WL (2011) Diagnostic radiograph based 3D bone reconstruction framework: application to the femur. Comput Med Imaging Graph 35(6):427–437
Amenta N, Choi S, Kolluri RK (2001) The power crust. In: Proceedings of the sixth ACM symposium on solid modeling and applications pp 249–266
Kies K, Benamrane N, Benyettou A (2005) 3D medical image segmentation and surface modeling using the power crust. Inf Technol J 4(4):377–381
Amenta N, Choi S, Kolluri RK (2001) The power crust, unions of balls, and the medial axis transform. Comput Geom Theory Appl 19(2):127–153
MURA - Musculoskeletal Radiographs. Available from: https://stanfordmlgroup.github.io/competitions/mura
LERA - Lower Extremity Radiographs. Available from: https://aimi.stanford.edu/lera-lower-extremity-radiographs
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition pp 770–778
Yu W, Chu C, Tannast M, Zheng G (2016) Fully automatic reconstruction of personalized 3D volumes of the proximal femur from 2D X-ray images. Int J Comput-Assist Radiol Surg 11(9):1673–1685
Funding
This paper is partially supported by Tongji University Sheng Feiyun College Student Science and Technology Innovation Practice Found and Student Innovation Training Program of Tongji University in 2021.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors Yunfei Ge, Qing Zhang, Yidong Shen, Yuantao Sun and Chongyang Huang declare that they have no conflict of interest.
Ethical approval
All procedures proposed in the paper including human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Ethics Committee of the First people’s Hospital of Yancheng ([2021]-(K-54)).
Informed consent
Informed consent has been obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ge, Y., Zhang, Q., Shen, Y. et al. A 3D reconstruction method based on multi-views of contours segmented with CNN-transformer for long bones. Int J CARS 17, 1891–1902 (2022). https://doi.org/10.1007/s11548-022-02701-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-022-02701-4