Abstract
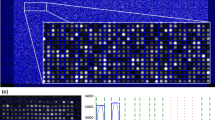
Image processing is an important stage of every microarray experiment. Reliability of this stage strongly influences the results of data analysis performed on extracted gene expressions. Multiple methods related to array recognition, spot segmentation and measurement extraction have emerged in this area over past several years. Currently there are various commercial and freeware packages available, which perform microarray image analysis. This paper attempts to review microarray image analysis as a whole and to make some experimental comparison of several computational schemes for signal segmentation and measurement extraction. Also we provide a detailed discussion of automated image quality control for use with microarray images.
Similar content being viewed by others
References
D.E. Bassett, M.B. Eisen, and M.S. Boguski, “Gene Expression Informatics—It's all in Your Mine,” Nature Genetics Supplement, 1999, p. 21, 48.
Y-X. Zhou, P. Kalocsai, J.-Y. Chen, and S. Shams, “Information Processing Issues and Solutions Associated with Microarray Technology,” in Microarray Biochip Technology, Schena (Ed.), Eaton Publishing, Massachusetts, 2000, pp. 167–200.
L.J. Heyer, S. Kruglyak, and S. Yooseph, “Exploring Expression Data: Identification and Analysis of Coexpressed Genes,” Genome Research, vol. 9, 1999, p. 1106.
Y.H. Yang, M.J. Buckley, S. Dudoit, and T.P. Speed, “Comparison of Methods for Image Analysis on cDNA Microarray Data,” Technical report, no. 584, Department of Statistics, University of California, Berkeley, 2000.
D. Bozinov and J. Rahnenf¨ uhrer, “Unsupervised Technique for Robust Target Separation and Analysis of Microarray Spots through Adaptive Pixel Clustering,” Bioinformatics, vol. 18, 2002, pp. 747–756.
R.A. Johnson and D.A. Wichern, Applied Multivariate Statistical Analysis. N.J.: Prentice Hall, 1998.
S.G. Hilsenbeck, W.E. Friedrichs, R. Schiff, P. O'Connell, R.K. Hansen, C.K. Osborne, and S.A.W. Fuqua, “Statistical Analysis of Array Expression Data as Applied to the Problem of Tamoxifen resistance,” Journal of the National Cancer Institute, vol. 91, 1999, p. 453.
Y. Chen, E.R. Dougherty, and M.L. Bittner, “Ratio-Based Decisions and the Quantitative Analysis of cDNA Microarray Images,” Journal of Biomedical Optics, vol. 2, no. 4, 1997.
R.O. Duda and P.E. Hart, Pattern Classification and Scene Analysis. J. Wiley, 1973.
R. Adams and L. Bischof, “Seeded Region Growing,” IEEE Transactions on Pattern Analysis and Machine Intelligence, no. 16, 1994.
K.R. Castleman, Digital Image Processing. Prentice Hall, 1996.
J. Brody, B. Williams, B. Wol, and S. Quake, “Significance and Statistical Errors in the Analysis of DNA Microarray Data,” PNAS, vol. 99, no. 20, 2002, pp. 12975–12978.
M.-L.T. Lee, F.C. Kuo, G.A. Whitmore, and J. Sklar, “Importance of Replication in Microarray Gene Expression Studies: Statistical Methods and Evidence from Repetitive cDNA Hybridizations,” PNAS,vol. 97, no. 18, 2000.
G.C. Tseng, M.K. Oh, L. Rohlin, J.C. Liao, and W.H. Wong, Issues in cDNAMicroarray Analysis: Quality Filtering, Channel Normalization, Models of Variations and Assessment of Gene Effects,” Nucleic Acids Res, vol. 29, 2001, pp. 2549–2557.
D.M. Mutch, A. Berger, R. Mansourian, A. Rytz, and M.A. Roberts, “The Limit Fold Change Model: A Practical Approach for Selecting Differentially Expressed Genes from Microarray Data,” BMC Bioinformatics, vol. 3, no. 17, 2002.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Petrov, A., Shams, S. Microarray Image Processing and Quality Control. The Journal of VLSI Signal Processing-Systems for Signal, Image, and Video Technology 38, 211–226 (2004). https://doi.org/10.1023/B:VLSI.0000042488.08307.ad
Published:
Issue Date:
DOI: https://doi.org/10.1023/B:VLSI.0000042488.08307.ad