Abstract
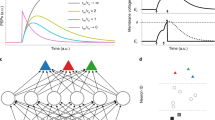
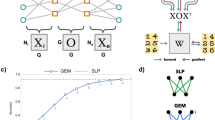
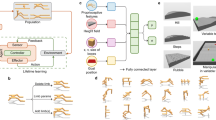
Although deep neural networks have seen great success in recent years through various changes in overall architectures and optimization strategies, their fundamental underlying design remains largely unchanged. Computational neuroscience may provide more biologically realistic models of neural processing mechanisms, but they are still high-level abstractions of empirical behaviour. Here we propose an evolvable neural unit (ENU) that can evolve individual somatic and synaptic compartment models of neurons in a scalable manner. We demonstrate that ENUs can evolve to mimic integrate-and-fire neurons and synaptic spike-timing-dependent plasticity. Furthermore, by constructing a network where an ENU takes the place of each synapse and neuron, we evolve an agent capable of learning to solve a T-maze environment task. This network independently discovers spiking dynamics and reinforcement-type learning rules, opening up a new path towards biologically inspired artificial intelligence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
A Code Ocean compute capsule, which contains a pre-built compute environment and the source code, is available at https://doi.org/10.24433/CO.1361267.v1.
References
Abbott, L. F. & Nelson, S. B. Synaptic plasticity: taming the beast. Nat. Neurosci. 3, 1178–1183 (2000).
Börgers, C. & Kopell, N. Synchronization in networks of excitatory and inhibitory neurons with sparse, random connectivity. Neural Comput. 15, 509–538 (2003).
Fell, J. & Axmacher, N. The role of phase synchronization in memory processes. Nat. Rev. Neurosci. 12, 105–118 (2011).
Hassabis, D., Kumaran, D., Summerfield, C. & Botvinick, M. Neuroscience-inspired artificial intelligence. Neuron 95, 245–258 (2017).
Spruston, N. Pyramidal neurons: dendritic structure and synaptic integration. Nat. Rev. Neurosci. 9, 206–221 (2008).
Sacramento, J., Costa, R. P., Bengio, Y. & Senn, W. Dendritic cortical microcircuits approximate the backpropagation algorithm. In Advances in Neural Information Processing Systems 8721–8732 (Curran Associates, 2018).
Jain, A. K., Mao, J. & Mohiuddin, K. M. Artificial neural networks: a tutorial. Computer 29, 31–44 (1996).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Catterall, W. A. Structure and function of voltage-gated ion channels. Ann. Rev. Biochem. 64, 493–531 (1995).
Flagel, S. B. et al. A selective role for dopamine in stimulus–reward learning. Nature 469, 53–57 (2011).
McCormick, D. A. Gaba as an inhibitory neurotransmitter in human cerebral cortex. J. Neurophysiol. 62, 1018–1027 (1989).
Levitan, I. B. & Kaczmarek, L. K. The Neuron: Cell and Molecular Biology (Oxford Univ. Press, 2015).
Nedergaard, M., Takano, T. & Hansen, A. J. Beyond the role of glutamate as a neurotransmitter. Nat. Rev. Neurosci. 3, 748–755 (2002).
Hilfiker, S. et al. Synapsins as regulators of neurotransmitter release. Philos. Trans. R. Soc. B 354, 269–279 (1999).
Hollenbeck, P. J. & Saxton, W. M. The axonal transport of mitochondria. J. Cell Sci. 118, 5411–5419 (2005).
Mountcastle, V. B. The columnar organization of the neocortex. Brain 120, 701–722 (1997).
Vale, R. D. The molecular motor toolbox for intracellular transport. Cell 112, 467–480 (2003).
Collingridge, G. L., Isaac, J. T. & Wang, Y. T. Receptor trafficking and synaptic plasticity. Nat. Rev. Neurosci. 5, 952–962 (2004).
Kepecs, A., Wang, X.-J. & Lisman, J. Bursting neurons signal input slope. J. Neurosci. 22, 9053–9062 (2002).
Abbott, L. F. Lapicque’s introduction of the integrate-and-fire model neuron (1907). Brain Res. Bull. 50, 303–304 (1999).
Hodgkin, A. L. & Huxley, A. F. A quantitative description of membrane current and its application to conduction and excitation in nerve. J. Physiol. 117, 500–544 (1952).
Hebb, D. O. The Organization of Behavior Vol. 65 (Wiley, 1949).
Maass, W. Networks of spiking neurons: the third generation of neural network models. Neural Networks 10, 1659–1671 (1997).
Bellec, G., Salaj, D., Subramoney, A., Legenstein, R. & Maass, W. Long short-term memory and learning-to-learn in networks of spiking neurons. In Advances in Neural Information Processing Systems 787–797 (Curran Associates, 2018).
Hornik, K. Approximation capabilities of multilayer feedforward networks. Neural Networks 4, 251–257 (1991).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Comput. 9, 1735–1780 (1997).
Cho, K. et al. Learning phrase representations using RNN encoder–decoder for statistical machine translation. Preprint at https://arxiv.org/abs/1406.1078 (2014).
Back, T. Evolutionary Algorithms in Theory and Practice: Evolution Strategies, Evolutionary Programming, Genetic Algorithms (Oxford Univ. Press, 1996).
Stanley, K. O., Clune, J., Lehman, J. & Miikkulainen, R. Designing neural networks through neuroevolution. Nature Mach. Intell. 1, 24–35 (2019).
Salimans, T., Ho, J., Chen, X., Sidor, S. & Sutskever, I. Evolution strategies as a scalable alternative to reinforcement learning. Preprint at https://arxiv.org/abs/1703.03864 (2017).
Frémaux, N. & Gerstner, W. Neuromodulated spike-timing-dependent plasticity, and theory of three-factor learning rules. Front. Neural Circuits 9, 85 (2016).
Mnih, V. et al. Human-level control through deep reinforcement learning. Nature 518, 529–533 (2015).
Hausknecht, M., Lehman, J., Miikkulainen, R. & Stone, P. A neuroevolution approach to general Atari game playing. IEEE Trans. Comput. Intell. AI Games 6, 355–366 (2014).
Igel, C. Neuroevolution for reinforcement learning using evolution strategies. In The 2003 Congress on Evolutionary Computation, 2003. CEC ‘03 Vol. 4, 2588–2595 (IEEE, 2003).
Bengio, S., Bengio, Y., Cloutier, J. & Gecsei, J. On the optimization of a synaptic learning rule. In Preprints Conf. Optimality in Artificial and Biological Neural Networks Vol. 2 (Univ. Texas, 1992).
Mouret, J.-B. & Tonelli, P. Artificial evolution of plastic neural networks: a few key concepts. In Growing Adaptive Machines 251–261 (Springer, 2014).
Risi, S. & Stanley, K. O. Indirectly encoding neural plasticity as a pattern of local rules. In International Conference on Simulation of Adaptive Behavior 533–543 (Springer, 2010).
Di Paolo, E. A. Evolving spike-timing-dependent plasticity for single-trial learning in robots. Philos. Trans. R. Soc. A 361, 2299–2319 (2003).
Carlson, K. D., Richert, M., Dutt, N. & Krichmar, J. L. Biologically plausible models of homeostasis and STDP: stability and learning in spiking neural networks. In The 2013 International Joint Conference on Neural Networks (IJCNN) 1–8 (IEEE, 2013).
Floreano, D., Epars, Y., Zufferey, J.-C. & Mattiussi, C. Evolution of spiking neural circuits in autonomous mobile robots. Int. J. Intell. Syst. 21, 1005–1024 (2006).
Rounds, E. L. et al. An evolutionary framework for replicating neurophysiological data with spiking neural networks. In International Conference on Parallel Problem Solving from Nature 537–547 (Springer, 2016).
Carlson, K. D., Nageswaran, J. M., Dutt, N. & Krichmar, J. L. An efficient automated parameter tuning framework for spiking neural networks. Front. Neurosci. 8, 10 (2014).
Buhry, L. et al. Automated parameter estimation of the Hodgkin–Huxley model using the differential evolution algorithm: application to neuromimetic analog integrated circuits. Neural Comput. 23, 2599–2625 (2011).
Venkadesh, S. et al. Evolving simple models of diverse intrinsic dynamics in hippocampal neuron types. Front. Neuroinformatics 12, 8 (2018).
Soltoggio, A., Durr, P., Mattiussi, C. & Floreano, D. Evolving neuromodulatory topologies for reinforcement learning-like problems. In 2007 IEEE Congress on Evolutionary Computation 2471–2478 (IEEE, 2007).
Blynel, J. & Floreano, D. Exploring the T-maze: evolving learning-like robot behaviors using CTRNNs. In Workshops on Applications of Evolutionary Computation 593–604 (Springer, 2003).
Doya, K. Metalearning and neuromodulation. Neural Networks 15, 495–506 (2002).
Soltoggio, A., Bullinaria, J. A., Mattiussi, C., Dürr, P. & Floreano, D. Evolutionary advantages of neuromodulated plasticity in dynamic, reward-based scenarios. In Proc. 11th International Conference on Artificial Life (Alife XI) 569–576 (MIT Press, 2008).
Back, T., Hoffmeister, F. & Schwefel, H.-P. A survey of evolution strategies. In Proc. 4th International Conference on Genetic Algorithms Vol. 2 (Morgan Kaufmann,1991).
Beyer, H.-G. & Schwefel, H.-P. Evolution strategies—a comprehensive introduction. Natural Comput. 1, 3–52 (2002).
Wierstra, D. et al. Natural evolution strategies. J. Mach. Learning Res. 15, 949–980 (2014).
Lehman, J., Chen, J., Clune, J. & Stanley, K. O. ES is more than just a traditional finite-difference approximator. In Proc. Genetic and Evolutionary Computation Conference 450–457 (ACM, 2018).
Sutskever, I., Martens, J., Dahl, G. & Hinton, G. On the importance of initialization and momentum in deep learning. In International Conference on Machine Learning 1139–1147 (JMLR, 2013).
Paszke, A. et al. Automatic Differentiation in PyTorch (Open Review, 2017).
Oliphant, T. E. A Guide to NumPy Vol. 1 (Trelgol, 2006).
Pawlak, V., Wickens, J. R., Kirkwood, A. & Kerr, J. N. Timing is not everything: neuromodulation opens the STDP gate. Front. Synaptic Neurosci. 2, 146 (2010).
Deacon, R. M. & Rawlins, J. N. P. T-maze alternation in the rodent. Nat. Protocols 1, 7–12 (2006).
Acknowledgements
We thank J. Kalafotovich, H. Bin Ko and R. Hormazabal for their review of the manuscript and related comments and discussions. This work was supported by the Institute for Information and Communications Technology Planning and Evaluation grant funded by the Korean government (MSIT) (no. 2017-0-01779, a machine learning and statistical inference framework for explainable artificial intelligence; no. 2019-0-01371, development of brain-inspired AI with human-like intelligence; and no. 2019-0-00079, Department of Artificial Intelligence, Korea University).
Author information
Authors and Affiliations
Contributions
P.B. conceived the network of ENUs and implemented related experiments. S.-W.L. discussed the results and supervised the research. P.B. and S.-W.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Machine Intelligence thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison and ablation study training progress.
Comparing standard GRUs, LSTMs and our proposed Evolvable Neural Unit (ENU). Generally the ENU consistently outperforms the other models in terms of final performance. Additionally to investigate the effect of the feedback connection from the output gate, we removed such connection in the No Feedback ENU (NFENU), showing the importance of this connection. In case of the Network of ENUs, we also ran additional experiments that shared the parameters between the neuron and synapse model (the SHAREDENUs-NN). It shows that having separate ENUs for both the synapses and neurons significantly improve performance, and that without such specialization the network fails to converge.
Extended Data Fig. 2 Mean input current vs Firing Frequency after evolving Integrate and Fire neurons.
shows that the evolved IAF model firing frequency pattern in response to the input current closely matches the underlying model it evolved to approximate.
Extended Data Fig. 3 Complex synaptic update rule example.
Comparing evolving an ENU for 1000 generations (left) vs 3000 generations (right). The ENU learned to approximate a complex neuromodulated STDP type learning rule. When the neurotransmitter is present at the input (NT) the rule follows a symmetric type STDP rule. However, when the NT signal is absent it follows completely different dynamics. It is maximum at a spike timing difference of around 0 and 10ms, while around 5ms the synaptic change is essentially disabled. This shows we do not require the manual derivation of a possibly complex exact mathematical function that explains the synaptic behaviour. Instead, ENUs can potentially evolve any type of complex arbitrary neuromodulated synaptic update learning rules when evolved within a larger complex network.
Extended Data Fig. 4 Double T-Maze evolved learning behaviour.
Double T-Maze evolved learning behaviour. Example of steps taking in the double T-maze environment by an evolved Network of ENUs. The agent can be seen to have successfully evolved to explore the environment to find and eat the initial poison (1). It then explores an alternative path to find non-poisonous food instead (2), indicating it has properly learned from a single example to associate the previous actions taken with a negative reward. Since food and poison can randomly change location, the agent goes back to the previous food location, but detects poison instead. As it previous obtained a negative reward with the action of eating the poison, it internally modified the synapse ENUs internal memory states to alter its behaviour, and successfully learned to turn around and find food in another part of the maze (3). It also evolved proper exploration behaviour if no food or poison is found in a section of the maze, successfully navigating to the other side (4).
Extended Data Fig. 5 Convergence analysis Ecomplex STDP and Double T-Maze experiment.
The ENU can be seen to generally converge faster in both experimental settings. In case of evolving a complex synaptic update rule (Complex STDP), the ENU significantly outperforms the other models. When the feedback connection is removed (NFENU), the performance also drops. This indicates the importance of the feedback connection, which was also observed in the previous standard STDP experiment in Fig. 4. In case of the Double T-Maze experiment, the ENU also converges faster with this feedback connection. The LSTM generally takes longer to converge compared to the GRU model, which could be explained by the fact that LSTMs are slightly more complex than GRUs. When the parameters are shared between the synapse and neuron ENUs, the network fails to converge (SHAREDENU). This was also observed in the standard T-Maze experiment, and further indicates the need for the specialization of the synaptic and neuronal behaviour.
Extended Data Fig. 6 Computation flow diagram of a Network of ENUs.
Shows a computation example with 4 ENU synapses and 2 ENU neurons, each having 3 channels. The sensory input neurons X are concatenated with all the ENU neurons H to get our input batch. A connection matrix is then applied that broadcasts (copies) the neurons’ output to each connected synapse ENU (1). On this resulting matrix we can then apply standard matrix multiplication and compute our synapse ENUs output in parallel (2). We can reshape this and sum along the first axis, as we have the same number of synapses for each neuron (3). This gives us the integrated synaptic input to each neuron ENU (4). Finally, we apply the neuron ENUs on this summated batch and obtain the output for each neuron in the ENU network (5). Each ENU has multiple outputs, so we have multiple channels that are processed by the ENU (the columns of each matrix), and we also have multiple neuron and synapse ENUs computed in parallel (the rows of each matrix).
Extended Data Fig. 7 IAF and STDP experimental setup.
For evolving the IAF ENU a single random graded potential is given as input (left). The goal of the ENU is then to approximate the underlying IAF rule. In case of evolving the STDP rule (right) multiple input channels are used: the graded input potential, the input spike, the neuromodulation signal (A-NT1) and the backpropagating spike. The target is then to output the modified graded input potential matching the STDP rule.
Extended Data Table 1 Final performance of Complex STDP approximation and Double T-Maze experiment.
Shows the mean and standard deviation of the performance in each environment over 30 trial runs. The ENU consistently outperforms the compared models (p<0.005). On the Double T-Maze Experiment, the LSTM and GRU model perform similarly (p=0.012), which was also observed in previous experiments. See also Fig. 5 for a more detailed convergence analysis.
Supplementary information
Rights and permissions
About this article
Cite this article
Bertens, P., Lee, SW. Network of evolvable neural units can learn synaptic learning rules and spiking dynamics. Nat Mach Intell 2, 791–799 (2020). https://doi.org/10.1038/s42256-020-00267-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-020-00267-x